Questions
- What is the Minimum Number of Substitutions for a Non-Informative Site?
- For a non-informative site, there can be any number of substitutions between the sequences without changing the phylogenetic relationships.
In other words, a non-informative site can have zero, one, or multiple substitutions and still have the same tree topology.
This is because non-informative sites do not provide any useful information for distinguishing between different evolutionary scenarios.
Therefore, the minimum number of substitutions for a non-informative site is zero, but it can be any number greater than zero without affecting the phylogenetic inference.
- For a non-informative site, there can be any number of substitutions between the sequences without changing the phylogenetic relationships.
- What is Unweighted Parsimony?
- Unweighted parsimony is a method of phylogenetic inference that assumes that all character state changes (i.e., substitutions, insertions, and deletions) have the same cost, regardless of the type or location of the change.
This means that each change is assigned a unit cost, and the goal of the analysis is to find the tree that requires the fewest total number of changes (i.e., the tree with the minimum number of steps). - The unweighted parsimony method is based on the principle of parsimony, which suggests that the simplest explanation or hypothesis that explains the data is usually the most likely to be correct.
In phylogenetics, this principle is used to infer the evolutionary relationships among a group of organisms based on their shared similarities and differences. - To apply unweighted parsimony, the first step is to construct a multiple sequence alignment of the DNA or protein sequences from the organisms of interest.
Next, the alignment is used to construct a matrix of character state changes, which is then used to construct a phylogenetic tree by searching for the tree with the minimum number of steps. - While unweighted parsimony is a simple and intuitive method of phylogenetic inference, it does have some limitations.
For example, it does not take into account the fact that some changes may be more likely to occur than others, or that different types of changes may have different costs. Therefore, more complex models of evolution, such as maximum likelihood and Bayesian inference, are often used in combination with parsimony to improve the accuracy of phylogenetic inference.
- Unweighted parsimony is a method of phylogenetic inference that assumes that all character state changes (i.e., substitutions, insertions, and deletions) have the same cost, regardless of the type or location of the change.
—————————————————————
IMPORTANTE Unweighted Parsimony:
Non importa quanti nucleotidi di ogni tipo ci sono, ma importa solo il numero di tipi differenti (da 1 solo fino ad un massimo di 4: A, T, G, C). Chiamiamo il numero di tipi di nucleotidi, allora il minimo numero di sostituzioni in un albero filogenetico sono:
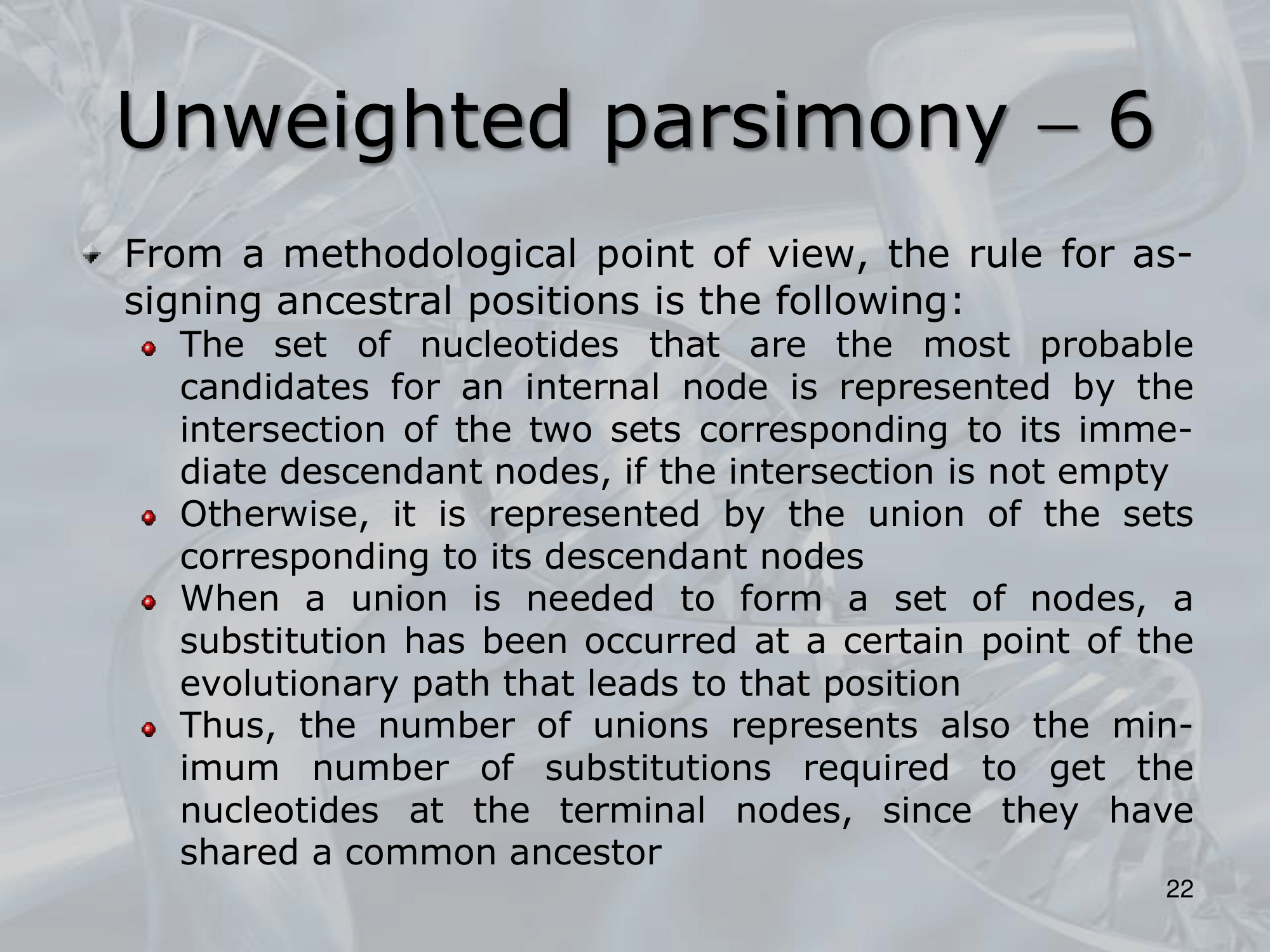
- Once non−informative sites are identified and discarded, the parsimony approach can be implemented in its simplest form:
- For each informative site, we consider the three possible trees.
- For each tree, a score is maintained that keeps track of the minimum number of substitutions required for each position.
- After considering all the informative sites, the tree (or the trees) which postulates the fewest number of substitutions is, by definition, the most parsimonious
- Computationally Challenging: The number of different unrooted trees grows exponentially with the number of sequences to be aligned, even having identified a small number of informative sites, this approach is inapplicable for more than 10 sequences.
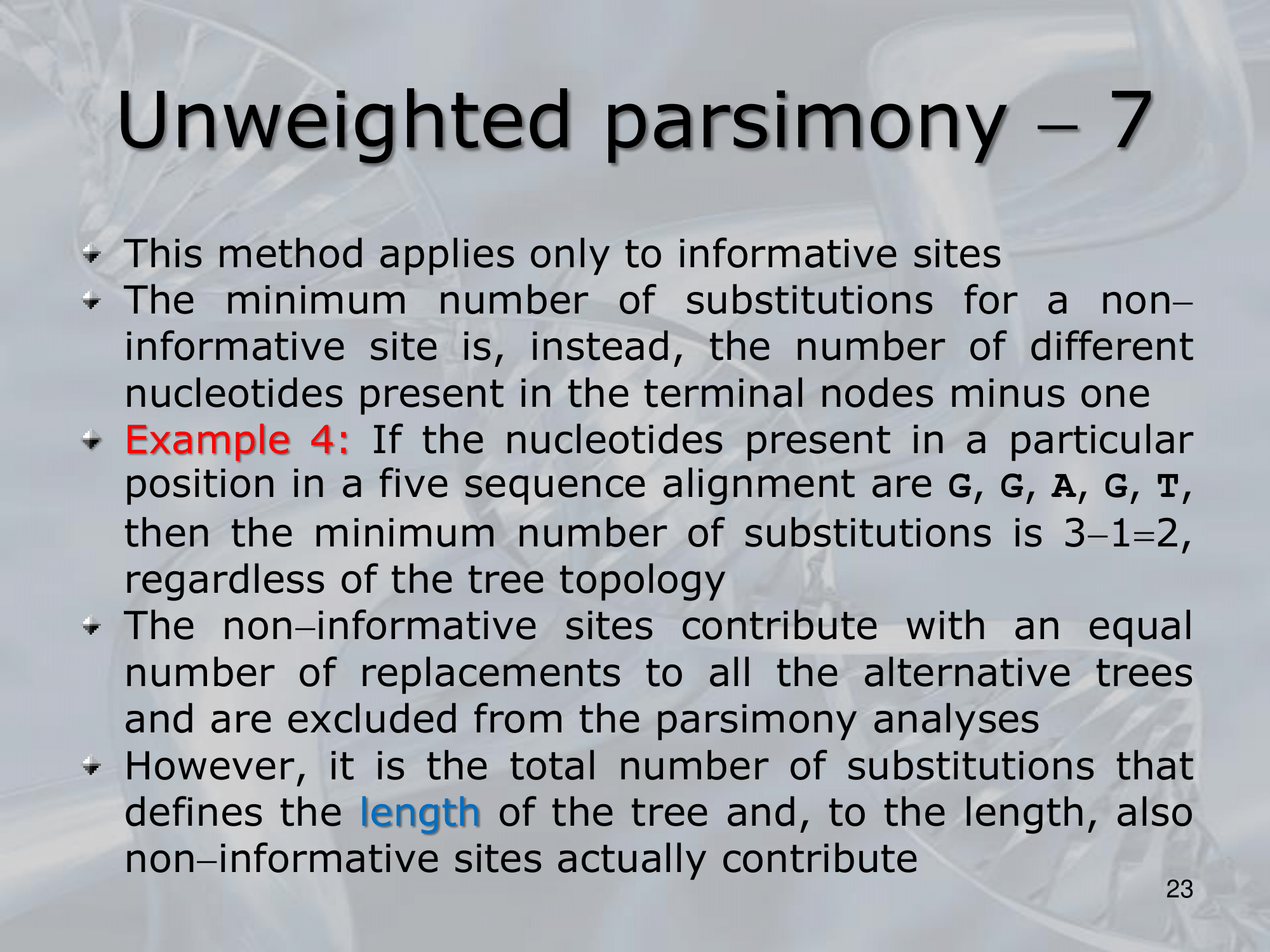
- This method applies only to informative sites
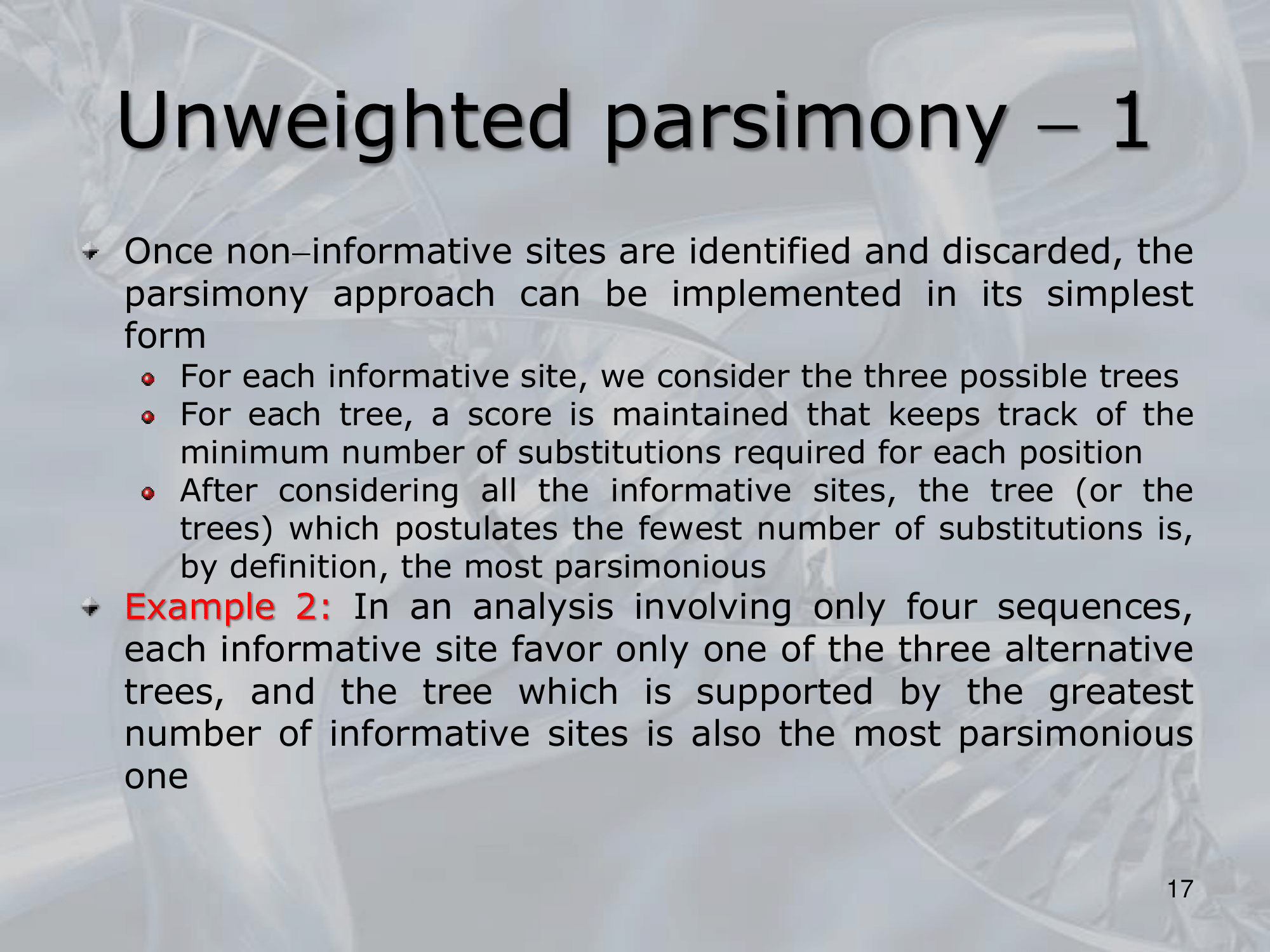
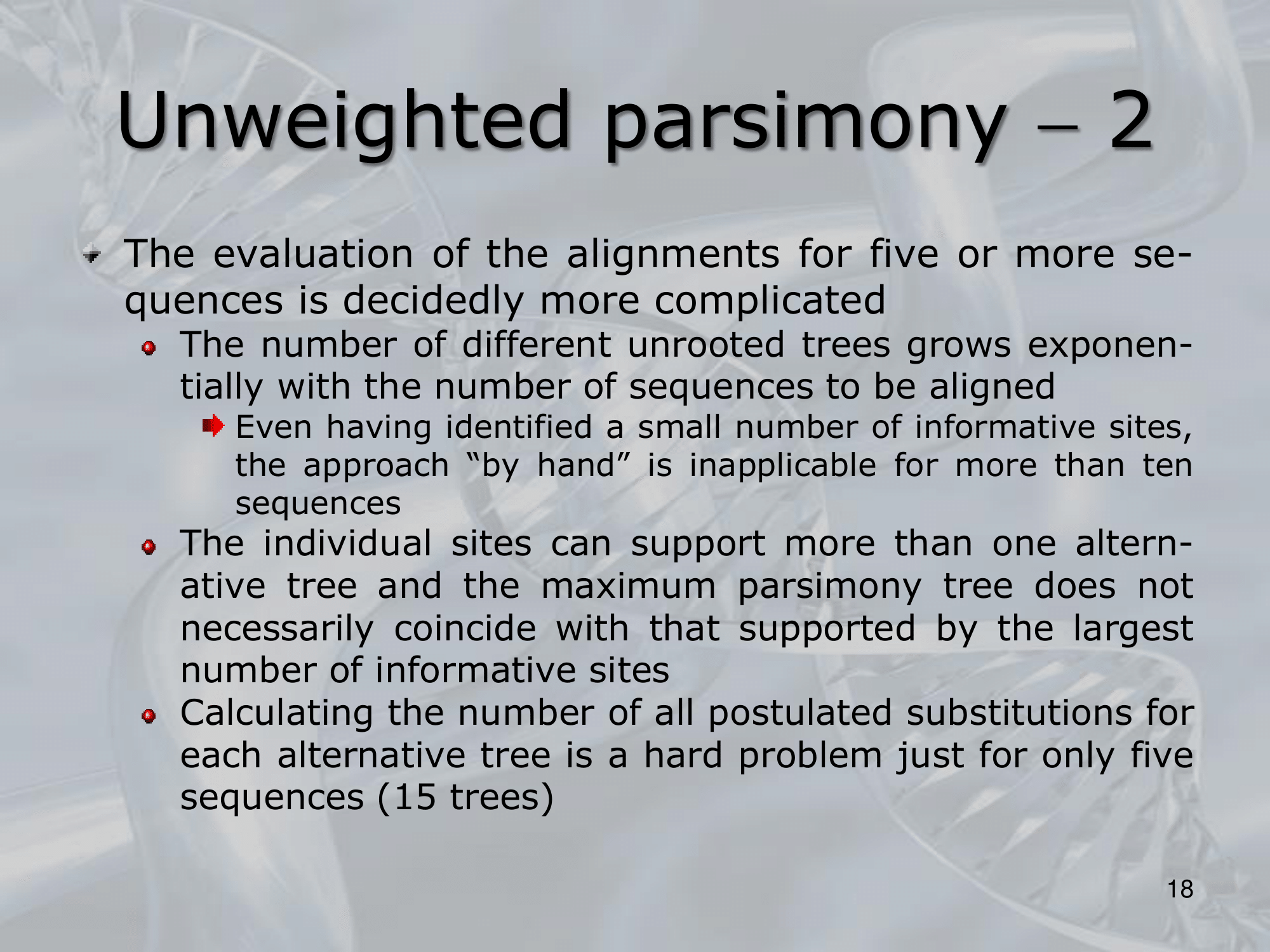
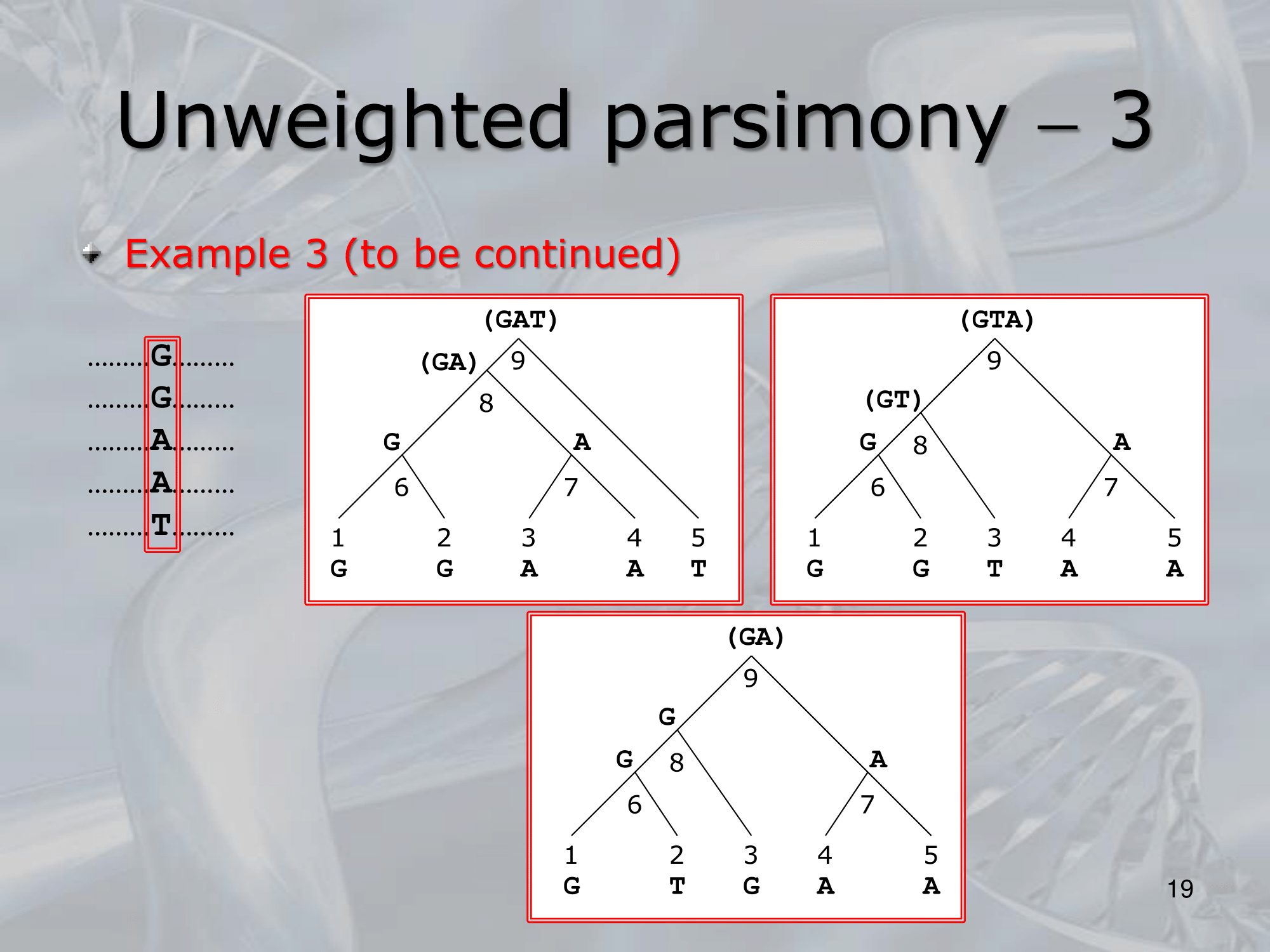
- What is the minimum number of substitution we can find? : The minimum number of substitutions for a non-informative site is, the number of different nucleotides present in the terminal nodes minus one ~Example: If the nucleotides present in a particular position in a five sequence alignment are G, G, A, G, T, then the minimum number of substitutions is , regardless of the tree topology
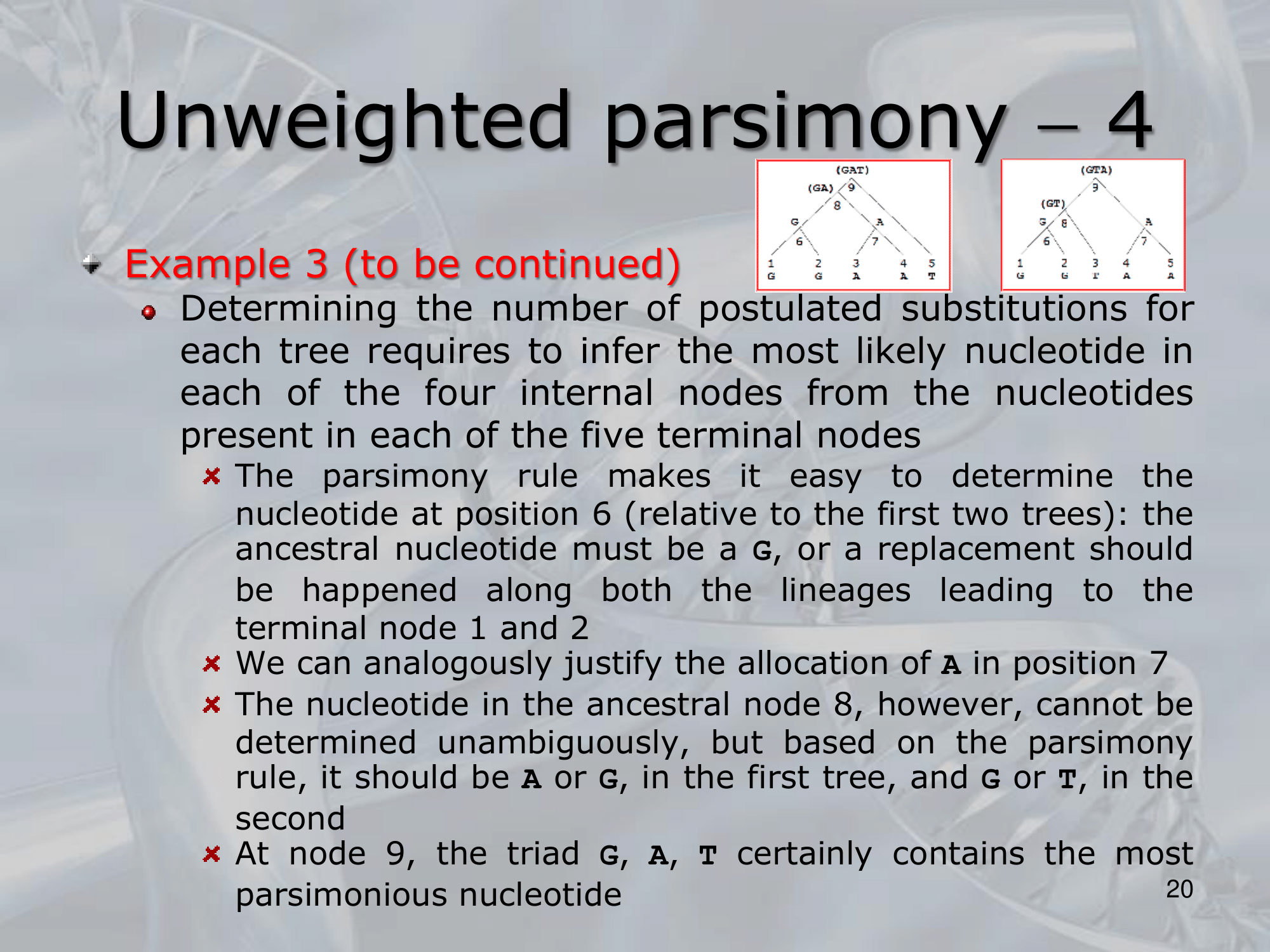
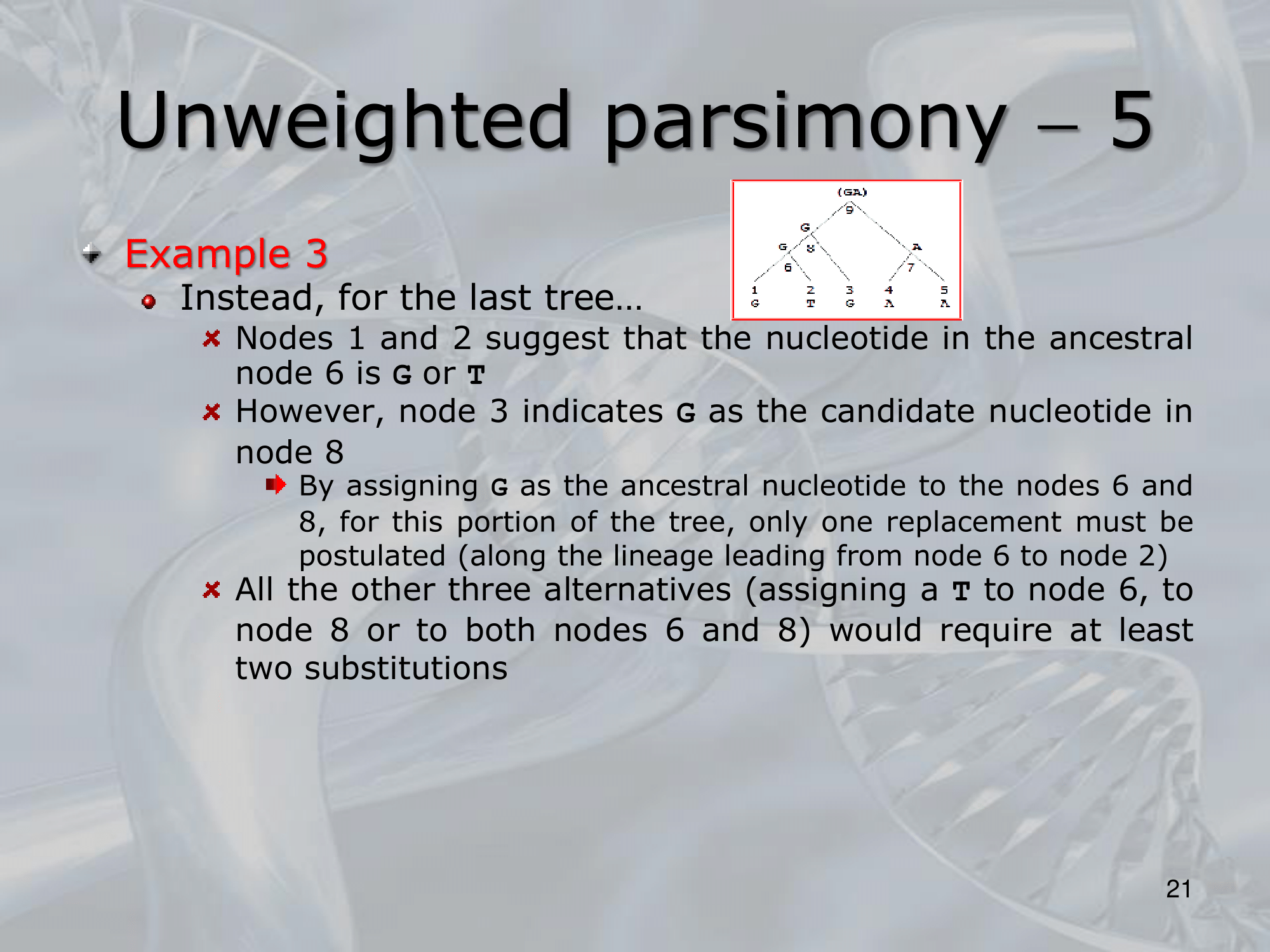
- ~Example: Here we can see three different way a single nucleotide changed using the rule of Unweighted Parsimony, all these three are equivalent because each of them count 3 total substitution (maximum parsimony rule)
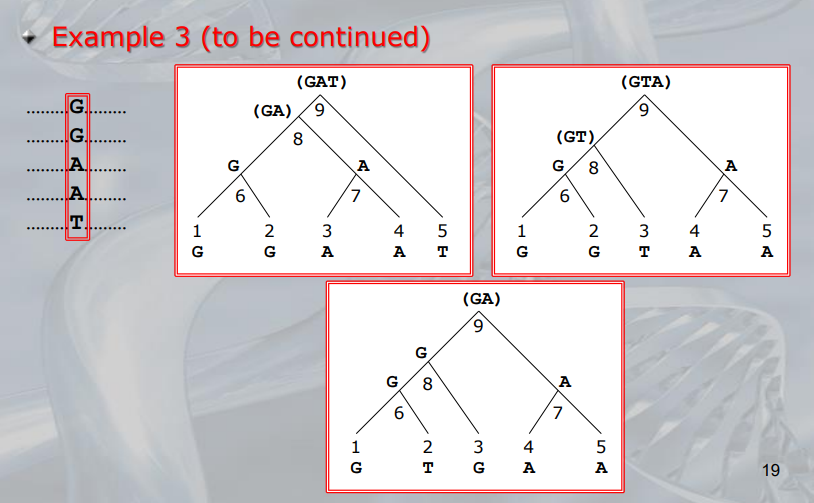
NOTE: The 3 trees are all equivalently parsimonious.
—————————————————————
Original Files