Questions
- What is there to say about the RNA Secondary Strucure Prediction?
- RNA secondary structure prediction is the process of determining the base pairing interactions between nucleotides within an RNA molecule, which is critical for understanding RNA structure and function.
==RNA secondary structure is important because it plays a crucial role in many biological processes, such as RNA splicing, translation, and gene expression regulation==. - RNA secondary structure prediction methods use computational algorithms to predict the most likely base pairing interactions between nucleotides within an RNA molecule.
==These methods typically use thermodynamic models, which calculate the stability of RNA structures based on the free energy of base pairing interactions==, and dynamic programming algorithms, which compute the most probable base pairs given the constraints of the thermodynamic model. - There are several RNA secondary structure prediction tools available, such as Mfold, RNAfold, and RNAstructure, which use different algorithms and thermodynamic models to predict RNA secondary structure.
Some of these tools can also predict RNA tertiary structure and RNA-protein interactions. - Despite significant progress in RNA secondary structure prediction, accurate prediction of RNA structures remains challenging due to the complex and diverse nature of RNA molecules.
The accuracy of RNA secondary structure prediction depends on the accuracy of the thermodynamic models, the choice of algorithm, and the quality of the input RNA sequence data.
Furthermore, some RNA molecules may form alternative or dynamic structures, which can be difficult to predict using current computational methods.
- RNA secondary structure prediction is the process of determining the base pairing interactions between nucleotides within an RNA molecule, which is critical for understanding RNA structure and function.
—————————————————————
IMPORTANTE
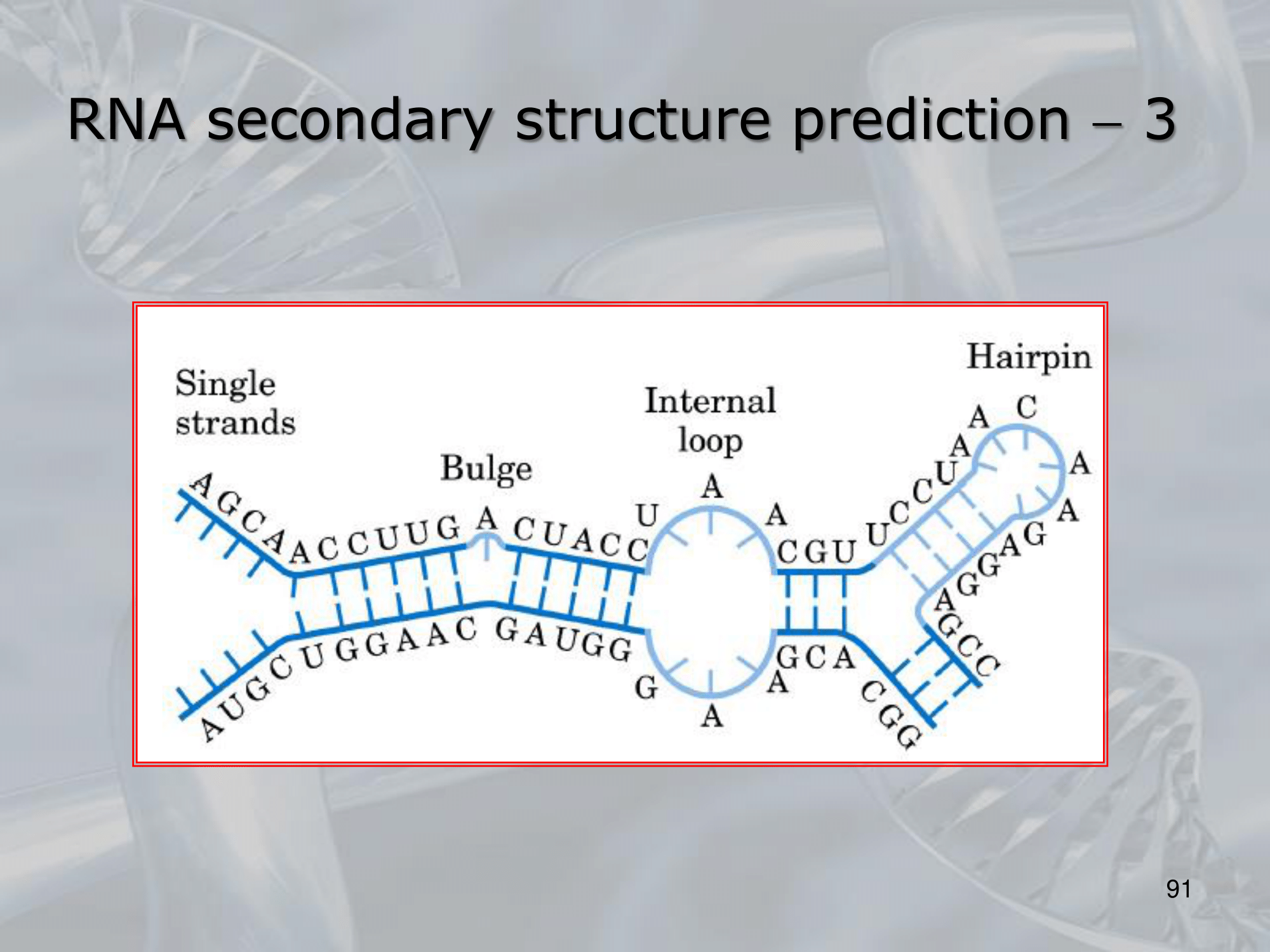
IMPORTANTE Unlike DNA, which usually assumes the well-known double helix conformation, the 2D stucture of a single-strand RNA is determined by the sequence of its nucleotides (like the protein is determined by its amminoacids). The RNA structure, however, is less complex than a protein structure. #IMPORTANTE ==In a SINGLE STRAND of RNA base paring can occur==, forming what are known as “secondary structures” #IMPORTANTE We can carattirze the 2D structure if we define where and which of the following secondary structure the RNA strand will have: Stems Hairpin Turn Bulge Loop
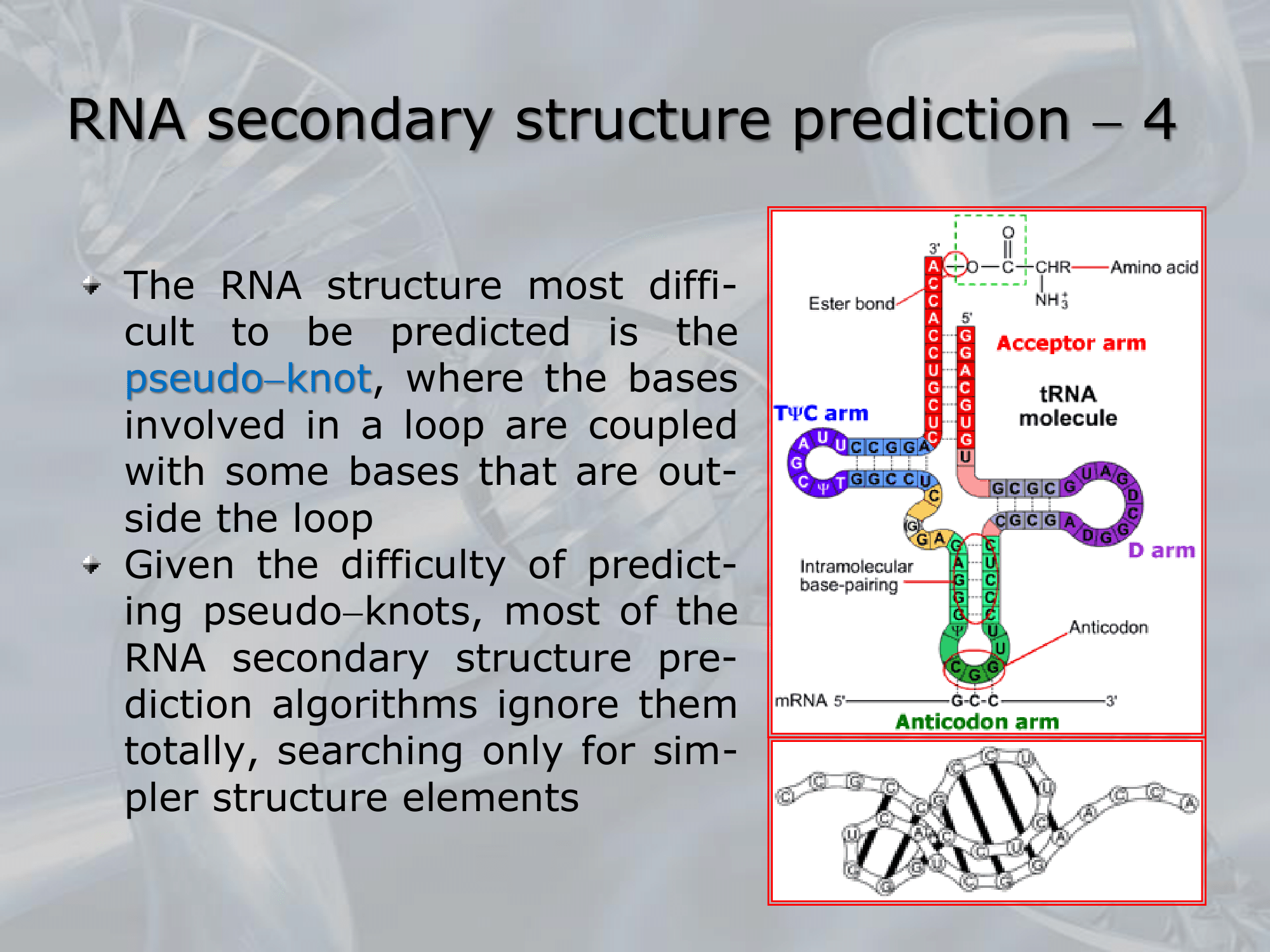
Pseudo-Knowts, when two different loops interact with each other folding the RNA strand:
Due to the difficutly of predicting pseudo-knowts, many algorithms for RNA secondary structure prediction ingore them all toghether
IMPORTANTE Understanding and correctly predicting the secondary structure formed by the single strand of DNA is important to understand gene regulation and expression of protein product. Since RNA is the product of gene-encoding DNA, and the traslation of RNA is one of the step to produce proteins.
IMPORTANTE In a recent descovery it was found that many RNAs strand have also a catalytic properties, these particular strands are called ribozymes, and they are involved in: Splicing of tRNA molecules. In the activity of ribosomes.
IMPORTANTE Also some viruses, such as HIV, are encoded in the form of RNA, so understaing its “folding” can help us fight such viruses. #IMPORTANTE Machine Learning approaches have been implemented, using as the cost function, the minimization of free energy of the final folded RNA, obtaining the most stable configuration of folded RNA.
—————————————————————
Slides with Notes