Questions
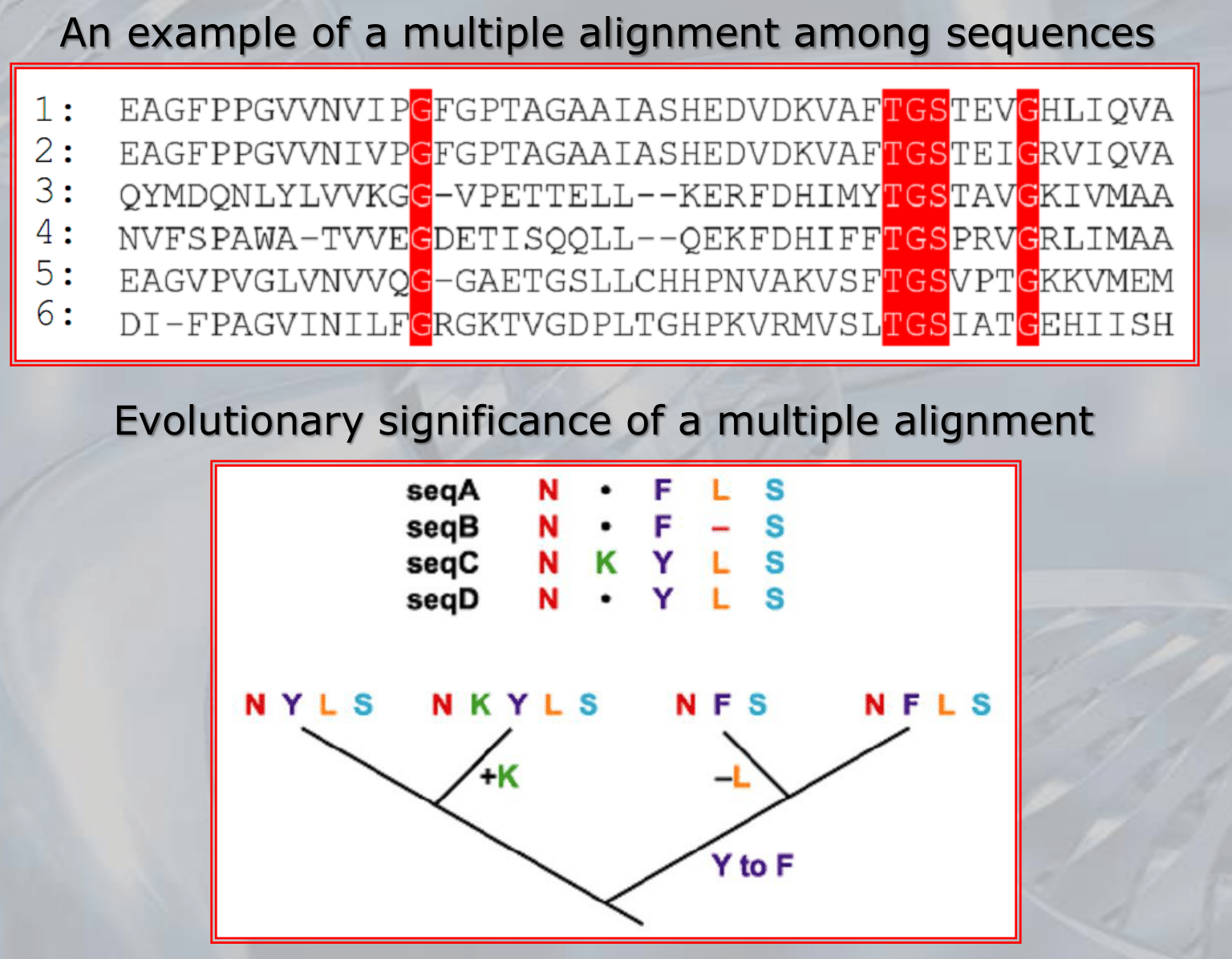
- What are Multiple Alignments?
- Multiple alignments refer to the process of aligning multiple sequences to identify the positions of similar or conserved residues in each sequence.
This is an important step in phylogenetic analysis as it allows for the comparison of nucleotide or amino acid sequences from different organisms.
Multiple alignments are used to construct phylogenetic trees and infer evolutionary relationships between organisms. - There are various algorithms and software programs available for multiple sequence alignment, including Clustal, MUSCLE, and T-Coffee.
The choice of method depends on the type and size of the dataset, the level of accuracy required, and the available computing resources.
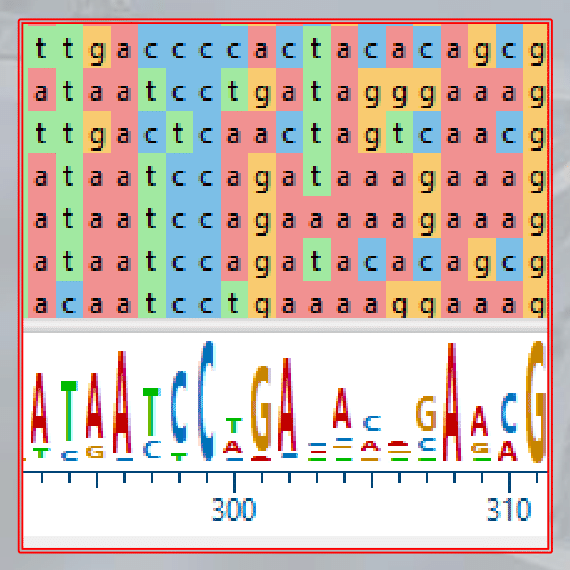
The resulting multiple alignment can be visualized as a matrix or alignment editor and is used to identify regions of conservation, gaps, and potential errors in the alignment.
- Multiple alignments refer to the process of aligning multiple sequences to identify the positions of similar or conserved residues in each sequence.
—————————————————————
IMPORTANTE
IMPORTANTE Multiple alignments: algoritmi che cercano di far risparmiare tempo quando si fanno confronti con più di 2 sequenze.
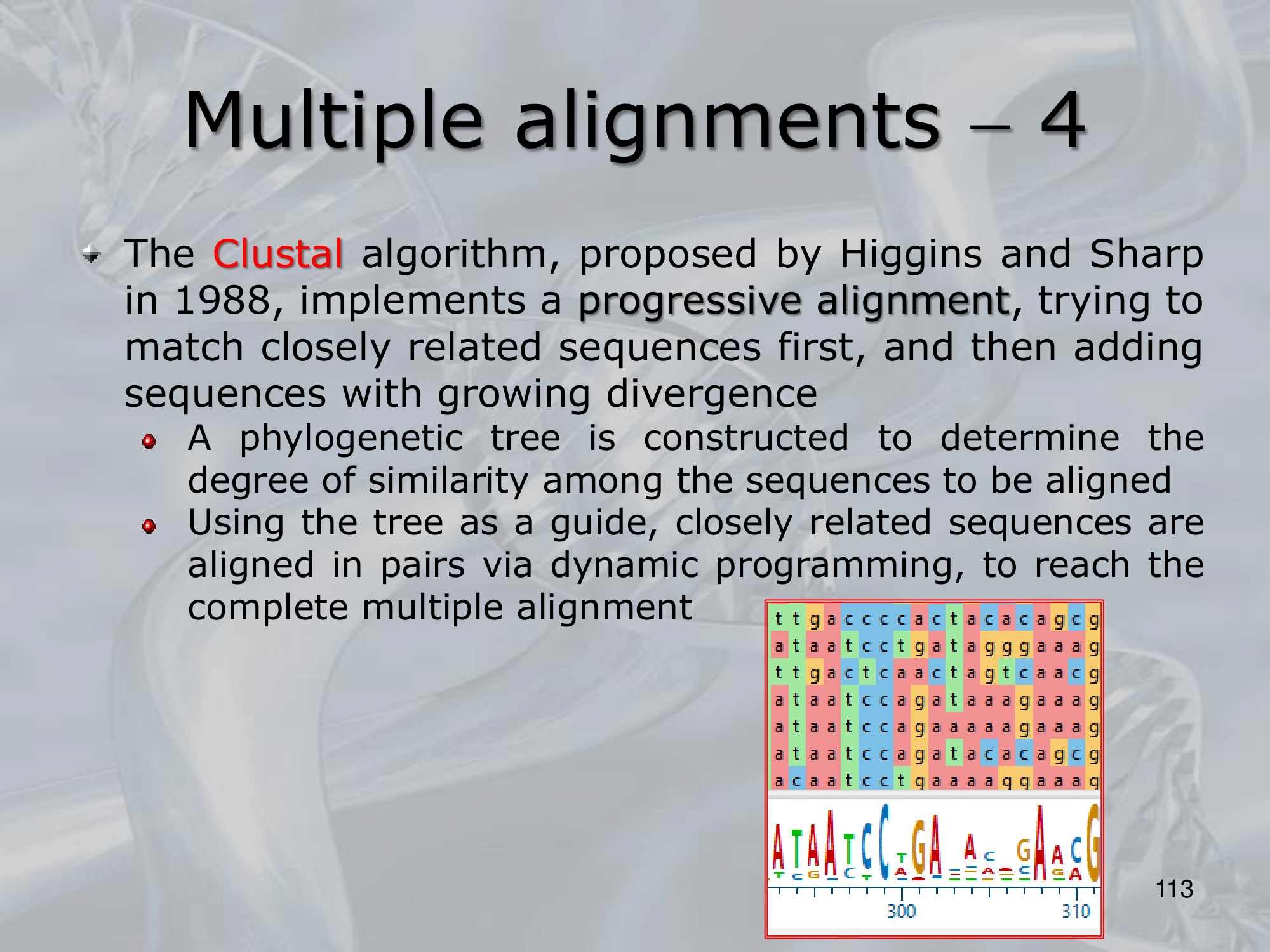
IMPORTANTE Clustal Algorithm for Multiple Alignments
- Tries to match closely related sequences first, then adds sequences with growing divergence
- A phylogenetic tree is constructed to determine the degree of similarity among the sequences to be aligned
- Usign the tree as a guide, closely related sequences are aligned in pairs via dynamic programming.
==The selection of ad hoc score matricies in fundamental to have a good clustal algorithm==.
~Ex.: Multiple Alignments
IMPORTANTE ClustalW The sequences are weighted according to their divergence from the pair of sequnces most closely realted to each other.
IMPORTANTE Another strategy for Multiple Alignments is to not penalize aligned gaps. #IMPORTANTE Other additional knowledge has to be added by hand, Clustal only watches for alignemnts of nucleotides and amino acids, impartially.
—————————————————————
Slides with Notes

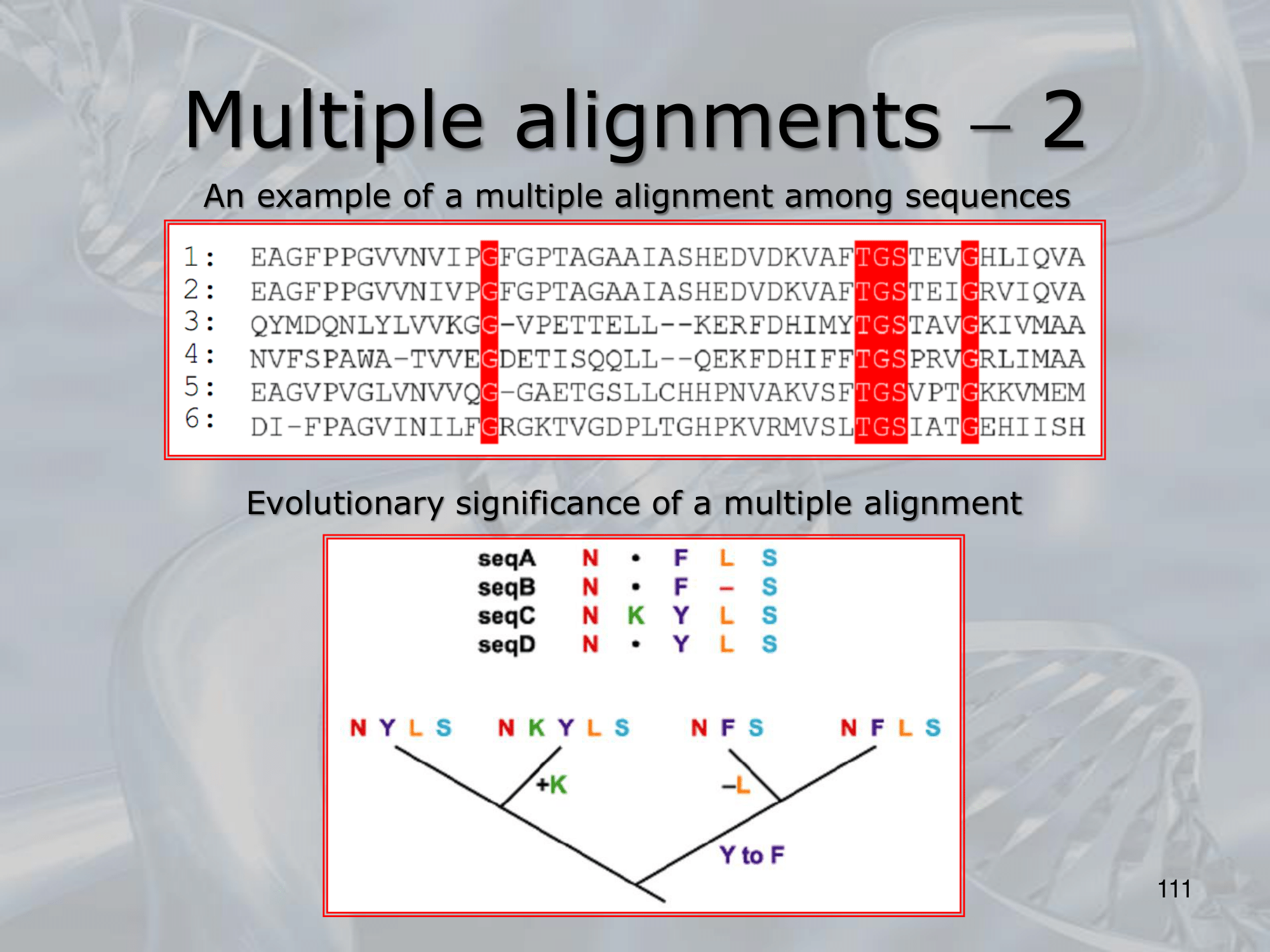


IMPORTANTE Clustal Algorithm for Multiple Alignments
- Tries to match closely related sequences first, then adds sequences with growing divergence
- A phylogenetic tree is constructed to determine the degree of similarity among the sequences to be aligned
- Usign the tree as a guide, closely related sequences are aligned in pairs via dynamic programming.
The selection of ad hoc score matricies in fundamental to have a good clustal algorithm.

IMPORTANTE ClustalW The sequences are weighted according to their divergence from the pair of sequnces most closely realted to each other.
IMPORTANTE Another strategy for Multiple Alignments is to not penalize aligned gaps. #IMPORTANTE Other additional knowledge has to be added by hand, Clustal only watches for alignemnts of nucleotides and amino acids, impartially.




IMPORTANTE Multiple alignments: algoritmi che cercano di far risparmiare tempo quando si fanno confronti con più di 2 sequenze.