Questions
- What is Alternative Splicing?
- ==Alternative splicing is a process in which different combinations of exons within a pre-mRNA transcript are spliced together to generate multiple mRNA isoforms from a single gene==.
This process is regulated by a complex network of splicing factors and can lead to the production of functionally distinct protein isoforms from a single gene. - The splicing of pre-mRNA transcripts is typically guided by splicing signals, which are specific nucleotide sequences located at the boundaries between exons and introns.
Alternative splicing can occur when these splicing signals are recognized in different ways, leading to the inclusion or exclusion of different exons in the mature mRNA transcript. - There are several types of alternative splicing events, including exon skipping, intron retention, alternative 5’ and 3’ splice site selection, and alternative promoter usage.
- Exon skipping occurs when one or more exons are excluded from the mature mRNA transcript, while intron retention occurs when one or more introns are included.
- Alternative 5’ and 3’ splice site selection occurs when different splice sites are used to join exons together, resulting in different combinations of exons in the mature mRNA.
- Alternative promoter usage occurs when different promoters are used to initiate transcription of a gene, leading to the inclusion of different exons in the mature mRNA.
- Alternative splicing plays a critical role in regulating gene expression and is thought to be a major contributor to the diversity of protein isoforms in eukaryotic organisms.
Dysregulation of alternative splicing has been linked to a variety of human diseases, including cancer and neurodegenerative disorders.
- ==Alternative splicing is a process in which different combinations of exons within a pre-mRNA transcript are spliced together to generate multiple mRNA isoforms from a single gene==.
—————————————————————
IMPORTANTE
IMPORTANTE hnRNA: In eukaryotes the primary transcript of RNA Polymerase II is the hnRNA (homogeneus RNA), which will be relocated into the cytoplasm, where it will be translated, undergo a series of changes (~ex.: the removal of introns)
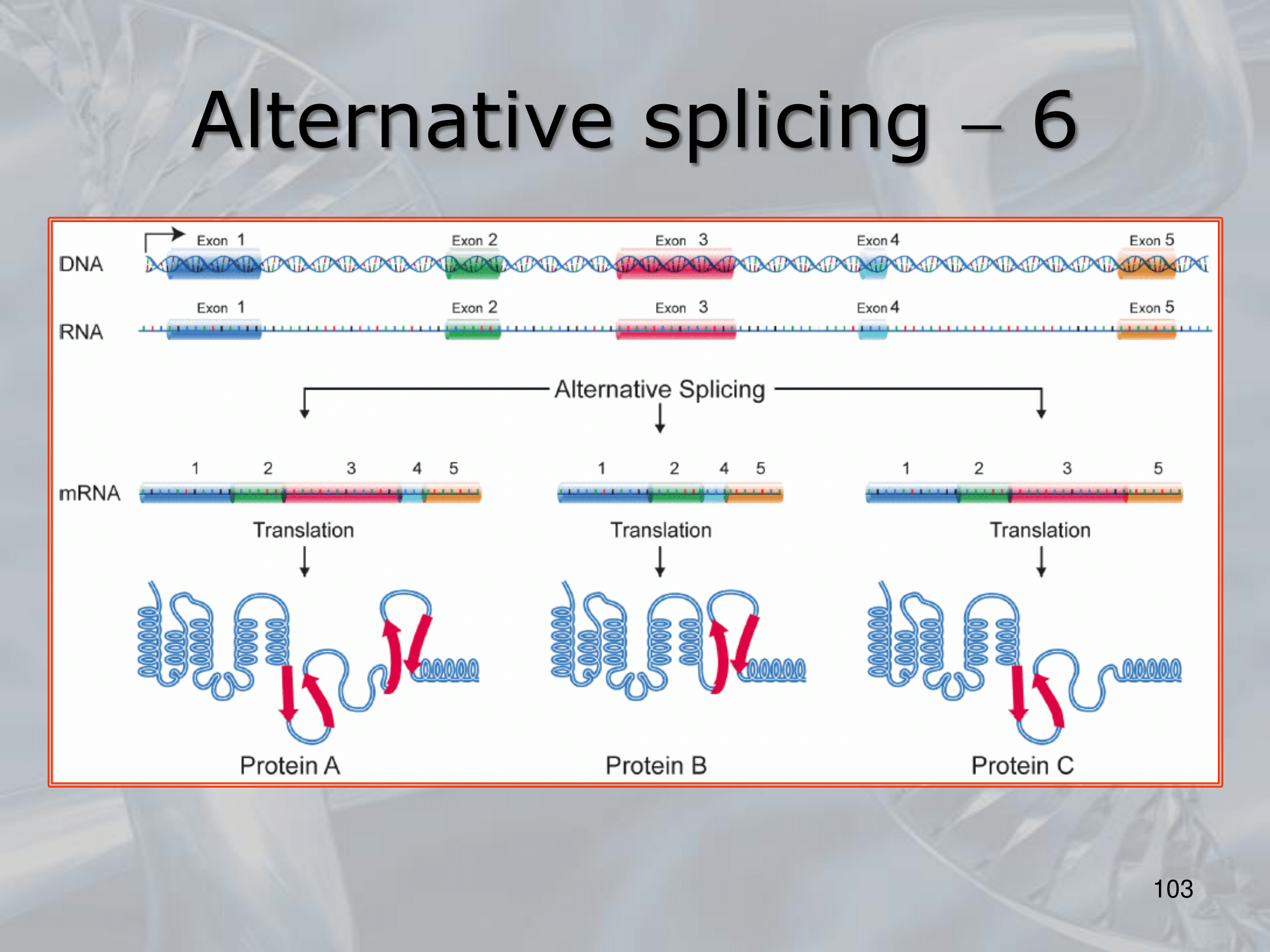
IMPORTANTE Alternative Splicing: The alternative splicing of a single mRNA, can generate (after translation) mature distinct protein isoforms.
Remember that:
- Many different eukarytic genes can be mapped into the same mRNA
- More than of eukaryotic genes, can generate multiple different mRNAs.
There are five different modes in which alternative splicing can occur:
- Exon Skipping: one exon is eliminated from the primary transcript (or preRNA or hnRNA).
- Mutually Exclusive Exons: only one out of two exons is kept.
- Alternative Cutting Site in 5’:
- Alternative Cutting Site in 3’:
- Intron Retention: an intron of the preRNA is kept in the mRNA.
Online Resource: Youtiube
IMPORTANTE In recent years it has become more clear that the “deregulation” of alternative splicing in genes can be the cause of the appearence of tumor cells.
—————————————————————
Slides with Notes

IMPORTANTE hnRNA: In eukaryotes the primary transcript of RNA Polymerase II is the hnRNA (homogeneus RNA), which will be relocated into the cytoplasm, where it will be translated, undergo a series of changes (~ex.: the removal of introns)
IMPORTANTE Alternative Splicing: The alternative splicing of a single mRNA, can generate (after translation) mature distinct protein isoforms.
Remember that:
- Many different eukarytic genes can be mapped into the same mRNA
- More than of eukaryotic genes, can generate multiple different mRNAs.
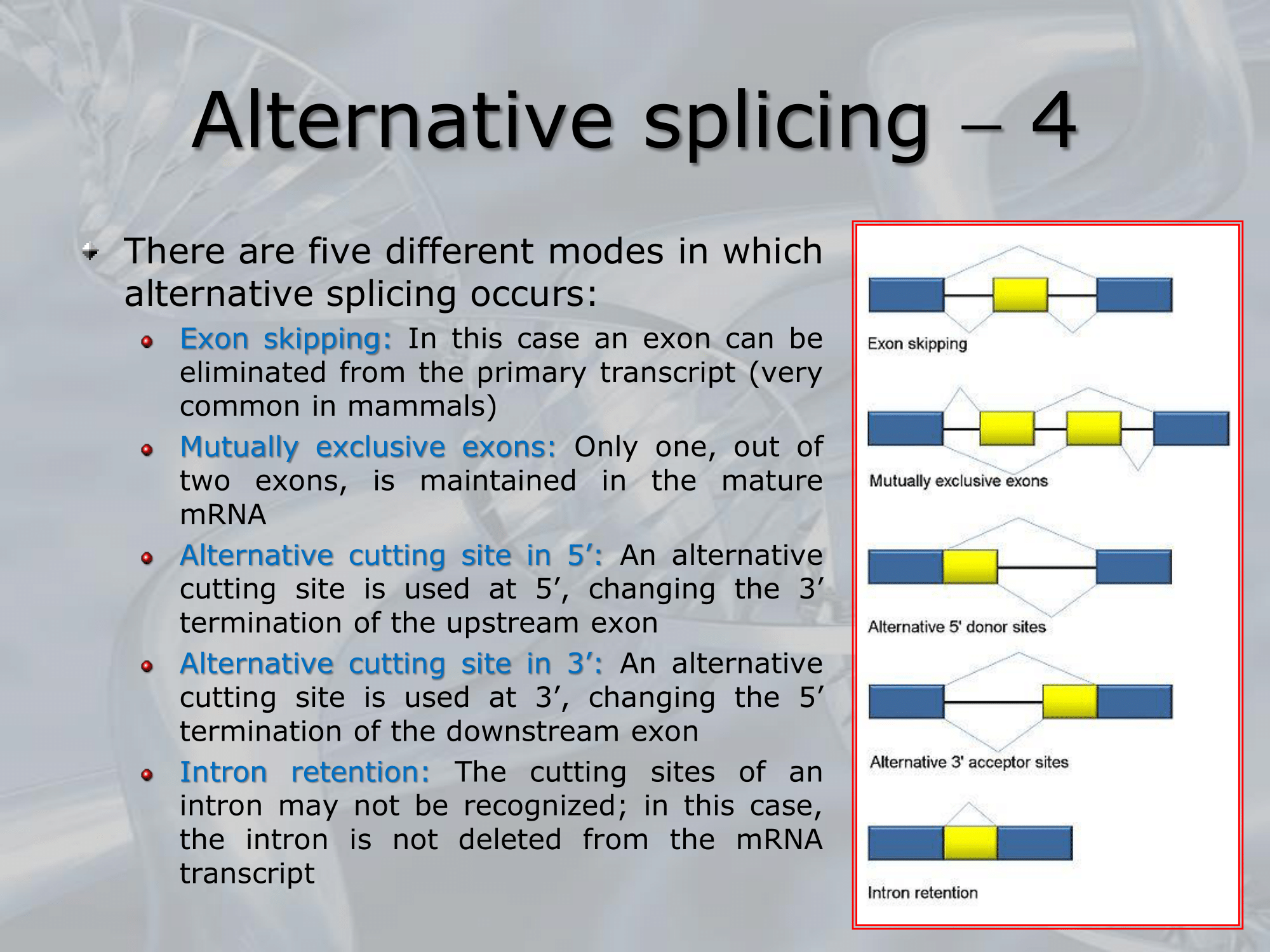
There are five different modes in which alternative splicing can occur:
- Exon Skipping: one exon is eliminated from the primary transcript (or preRNA or hnRNA).
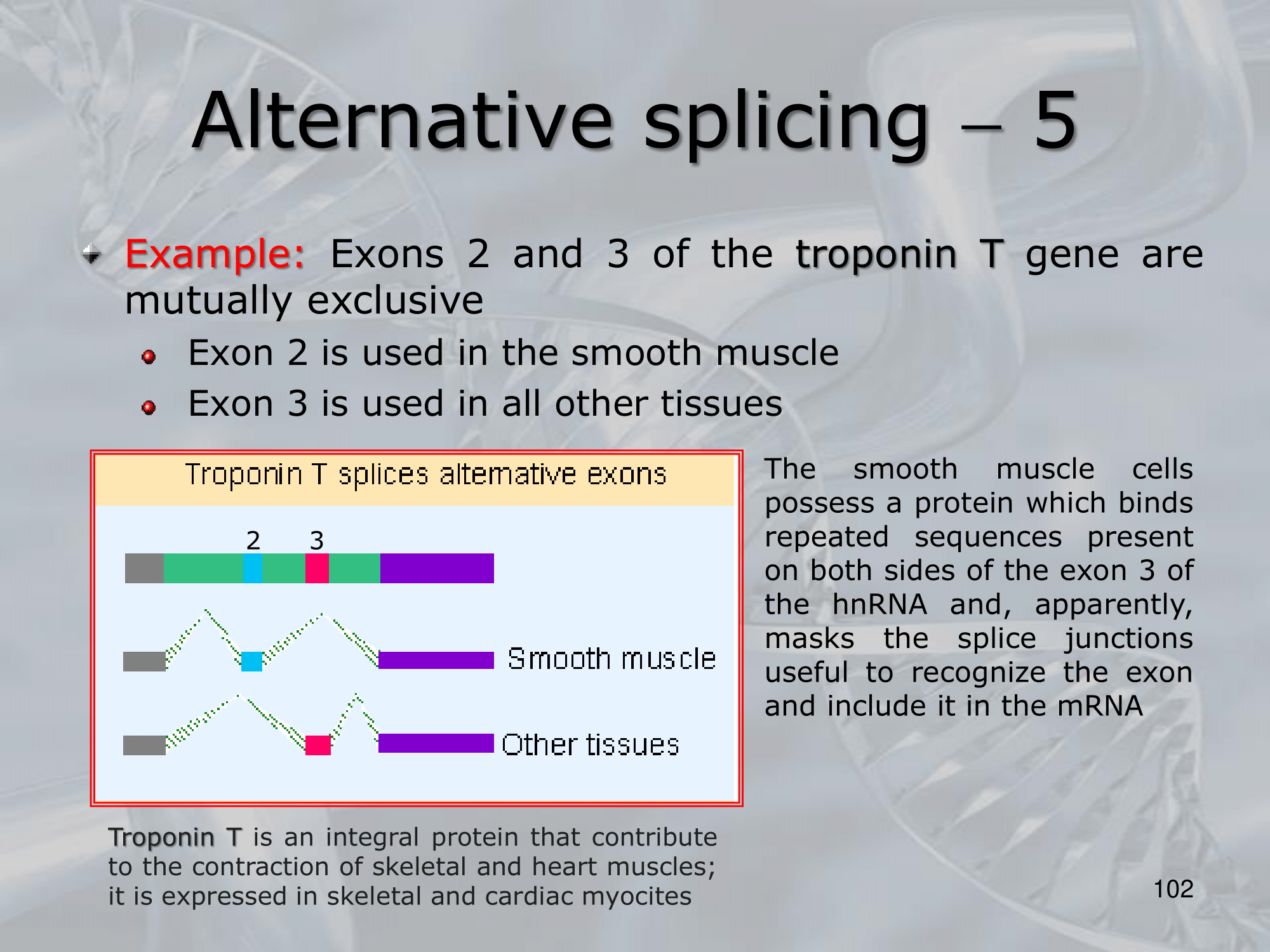
- Mutually Exclusive Exons: only one out of two exons is kept.
- Alternative Cutting Site in 5’:
- Alternative Cutting Site in 3’:
- Intron Retention: an intron of the preRNA is kept in the mRNA.
Online Resource: Youtiube





IMPORTANTE In recent years it has become more clear that the “deregulation” of alternative splicing in genes can be the cause of the appearence of tumor cells.