Questions
- How do Protein Change?
- Proteins can change through a variety of mechanisms, including mutations, gene duplication, gene fusion, and horizontal gene transfer. Some of the most common mechanisms that lead to changes in proteins include:
- Point mutations: This is a change in a single nucleotide that leads to a change in the amino acid sequence of the protein.
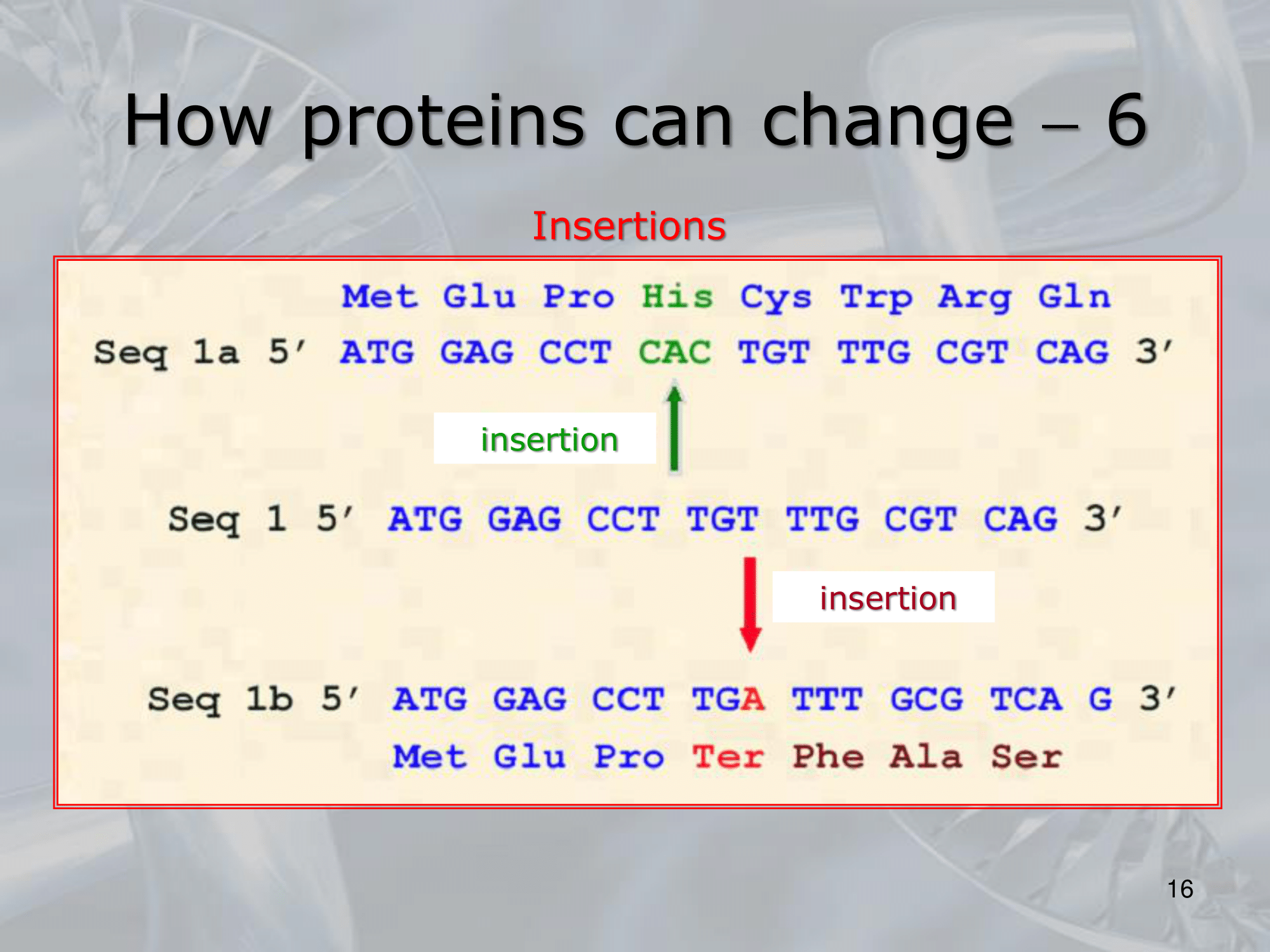
These mutations can be either synonymous, meaning they do not change the amino acid sequence, or non-synonymous, meaning they result in a change in the amino acid sequence. - Insertions and deletions: These are changes in the DNA sequence that result in the addition or removal of one or more nucleotides.
These changes can lead to frameshift mutations that alter the reading frame and often result in a completely different protein. - Gene duplication and divergence: Gene duplication can lead to the creation of new protein-coding genes, which can then evolve independently through the accumulation of point mutations, insertions, and deletions.
Over time, these changes can result in significant differences in the amino acid sequence and function of the duplicated proteins. - Gene fusion: Gene fusion occurs when two or more genes are joined together to form a hybrid gene.
This can lead to the formation of new proteins with novel functions. - Horizontal gene transfer: Horizontal gene transfer occurs when genes are transferred between different organisms, often through mechanisms such as viral infection or bacterial conjugation.
This can lead to the acquisition of new genes and proteins that may provide a selective advantage.
- Point mutations: This is a change in a single nucleotide that leads to a change in the amino acid sequence of the protein.
- Overall, changes in proteins can have significant impacts on the structure and function of the protein, as well as on the organism as a whole.
Understanding the mechanisms that lead to changes in proteins is important for understanding the evolution of life and for developing new approaches to treat and prevent disease.
- Proteins can change through a variety of mechanisms, including mutations, gene duplication, gene fusion, and horizontal gene transfer. Some of the most common mechanisms that lead to changes in proteins include:
- What is Adaptive Convergence?
- Adaptive convergence is a phenomenon in evolutionary biology where unrelated organisms evolve similar traits or features in response to similar environmental pressures or selective forces.
This convergence occurs because the selective pressure favors certain adaptations that increase fitness or survival in the environment, leading to the independent evolution of similar traits in different lineages. - ==For example, the wings of birds and bats are not homologous, meaning they do not have a common evolutionary origin.
However, both birds and bats evolved wings independently as a response to the selective pressure of flight==.
This is an example of adaptive convergence, where two unrelated groups of organisms evolved similar traits due to similar ecological pressures. - Adaptive convergence can also occur at the molecular level, where different genes or proteins in different organisms evolve similar amino acid sequences or functional domains as a response to similar selective pressures.
This convergence can be detected using molecular phylogenetics and comparative genomics approaches, and can provide insights into the adaptive evolution of different lineages in response to changing environments.
- Adaptive convergence is a phenomenon in evolutionary biology where unrelated organisms evolve similar traits or features in response to similar environmental pressures or selective forces.
- What is the Difference between Similarity and Homology, which one of them is a unit we can Measure?
- Similarity and homology are both concepts used in molecular biology and bioinformatics to describe the relationship between sequences of genes, proteins, or other biomolecules.
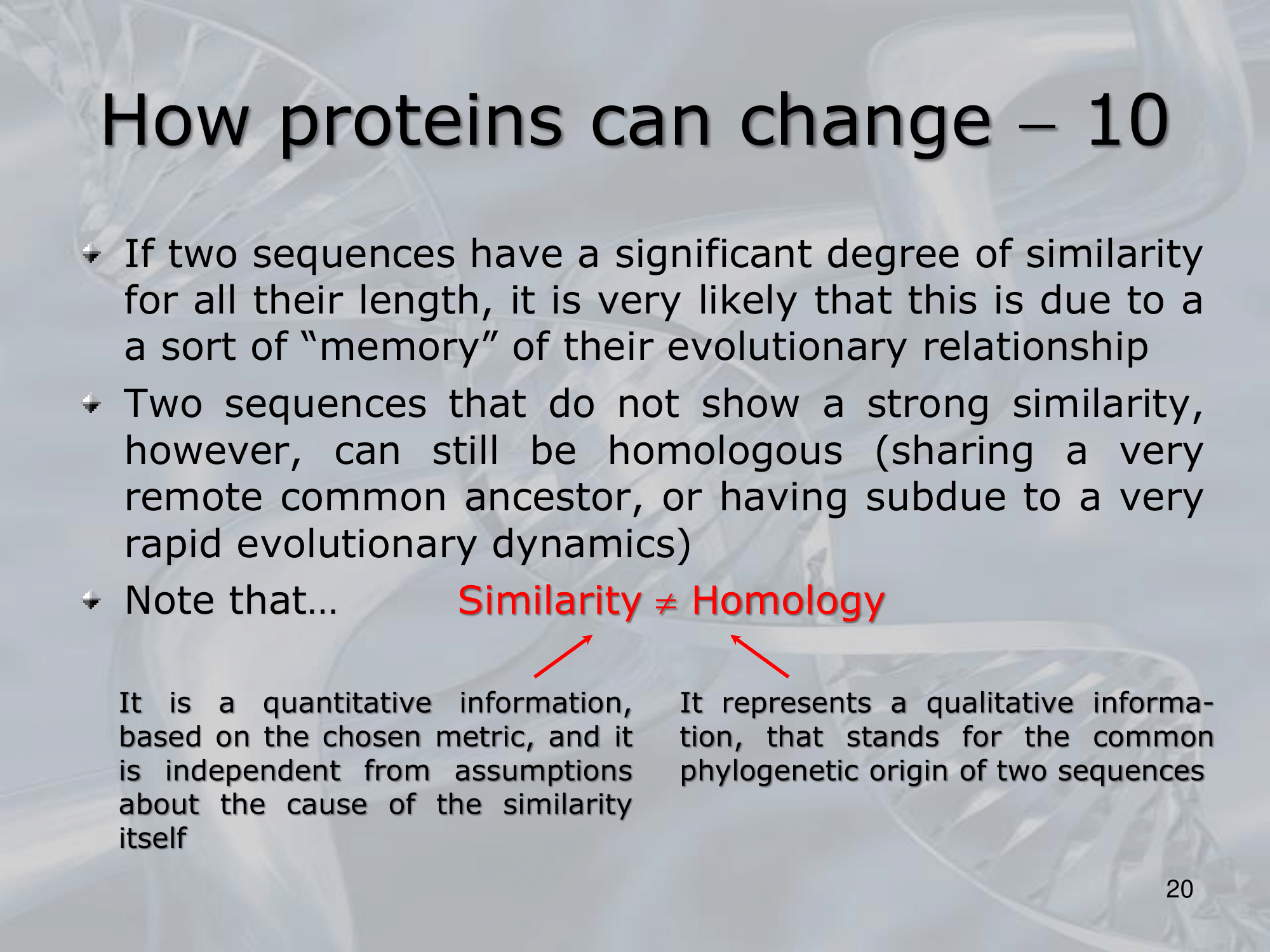
- Similarity refers to the degree of resemblance between two sequences, usually measured as a percentage of identical amino acids or nucleotides in a given alignment.
==Similarity can arise due to chance or convergent evolution, and does not necessarily imply a common evolutionary origin or function==. - Homology, on the other hand, refers to the presence of shared ancestry between two sequences, implying a common evolutionary origin and often a similar or related function.
Homology can be inferred by comparing the sequence similarity, but can also be supported by other evidence such as conserved gene synteny or functional data.
- Similarity refers to the degree of resemblance between two sequences, usually measured as a percentage of identical amino acids or nucleotides in a given alignment.
- While similarity can be quantified as a percentage of identical residues or nucleotides, homology is not a unit of measurement per se, but rather a qualitative description of the evolutionary relationship between sequences.
- In summary, similarity and homology are related but distinct concepts in molecular biology, with similarity being a quantitative measure of sequence resemblance and homology indicating a qualitative inference of shared ancestry and function.
- Similarity and homology are both concepts used in molecular biology and bioinformatics to describe the relationship between sequences of genes, proteins, or other biomolecules.
—————————————————————
IMPORTANTE
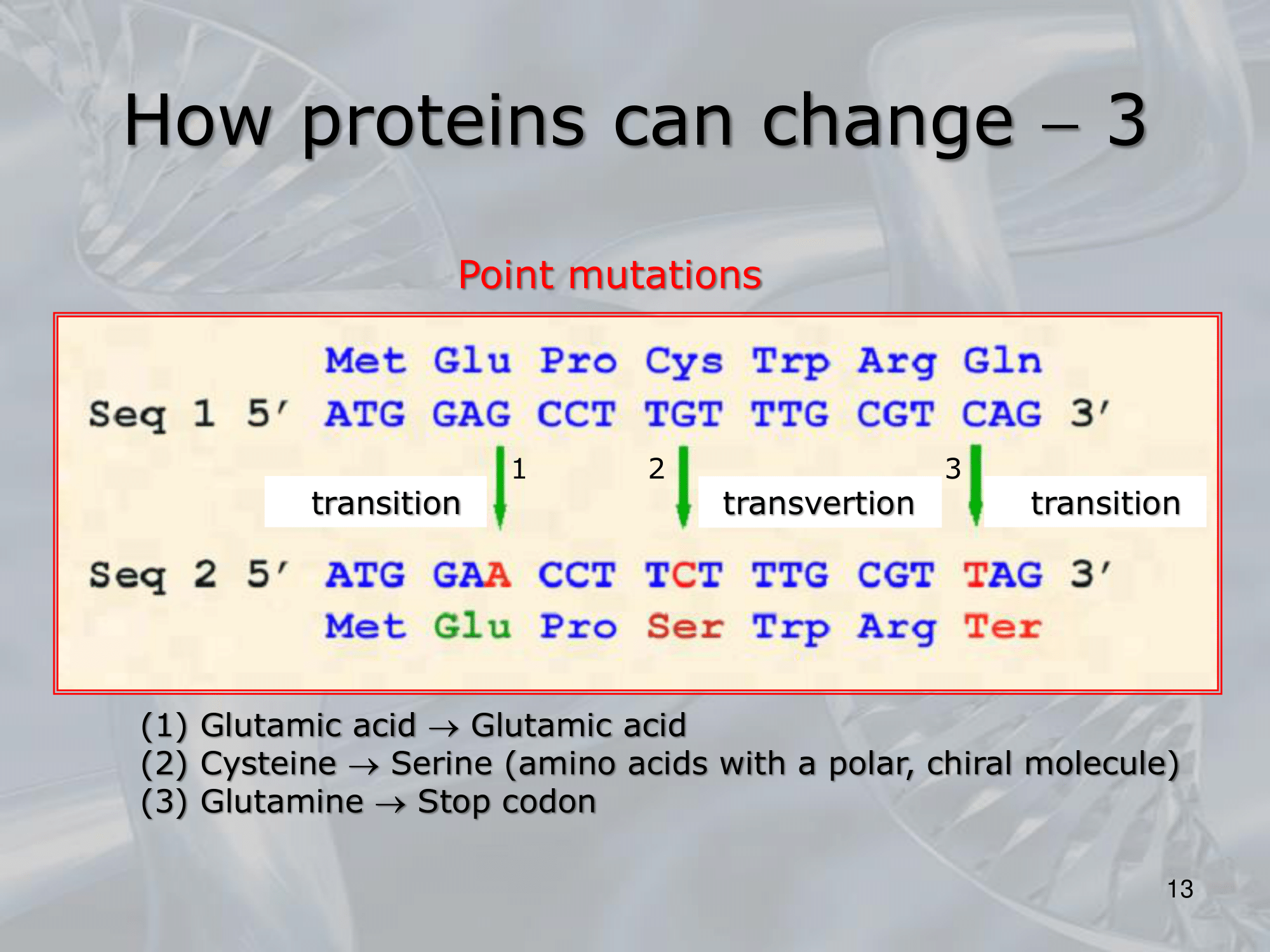
IMPORTANTE Two Types of Point Mutation or Substitution:
- Transition: if a Purine (G,A) changes into another Purine (A,G), same with Phyrimidine
- Traslation: if a Purine (G,A) becomes a Phyrimidine (C, T), or vice-versa
IMPORTANTE Il DNA di una specie è soggetto a diverse mutazioni:
- Sostituzione Puntuale: Una singola base cambia
- Inserzione: Viene aggiunta una nuova base.
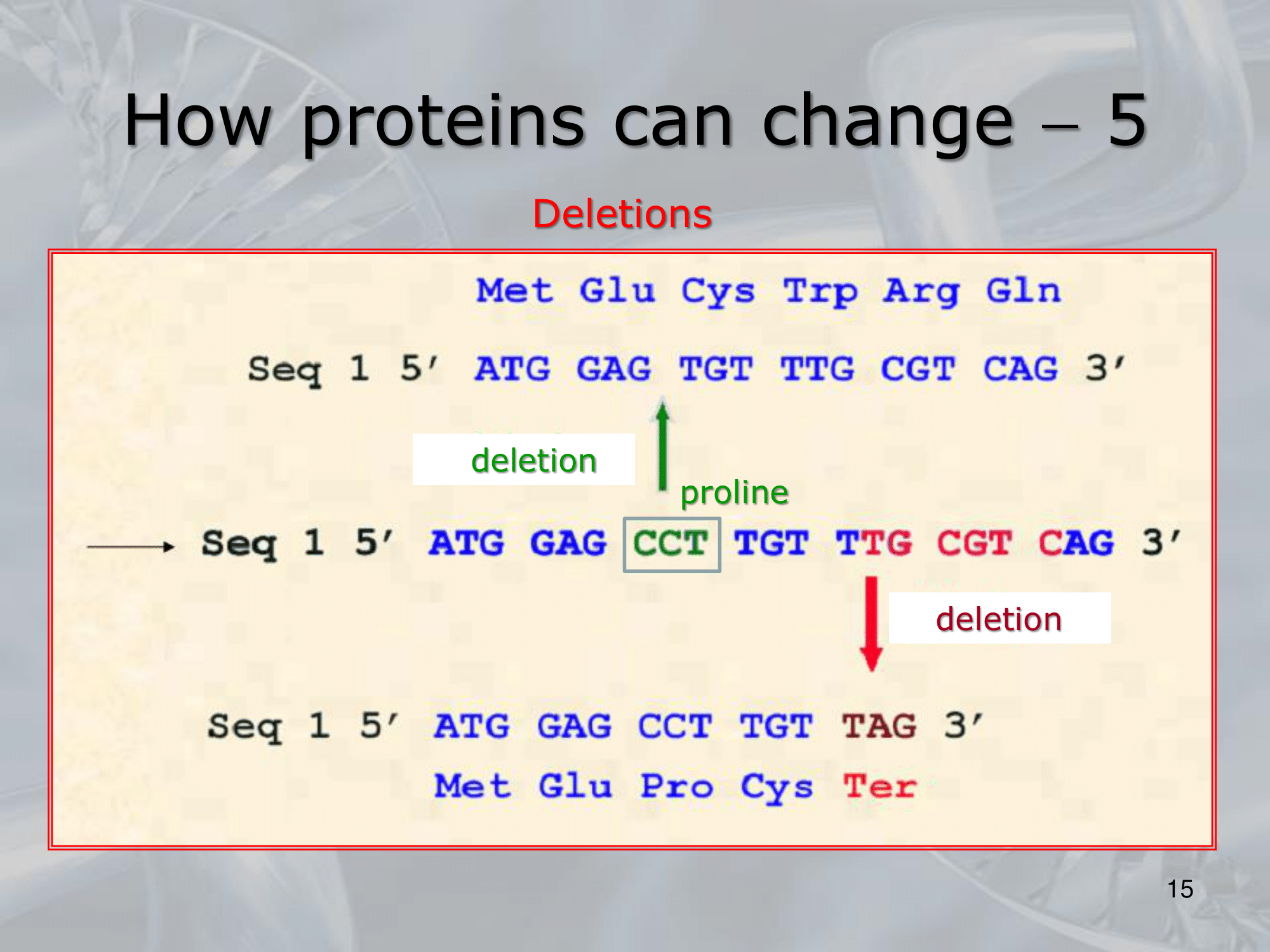
- Cancellazione: Viene tolta una base.
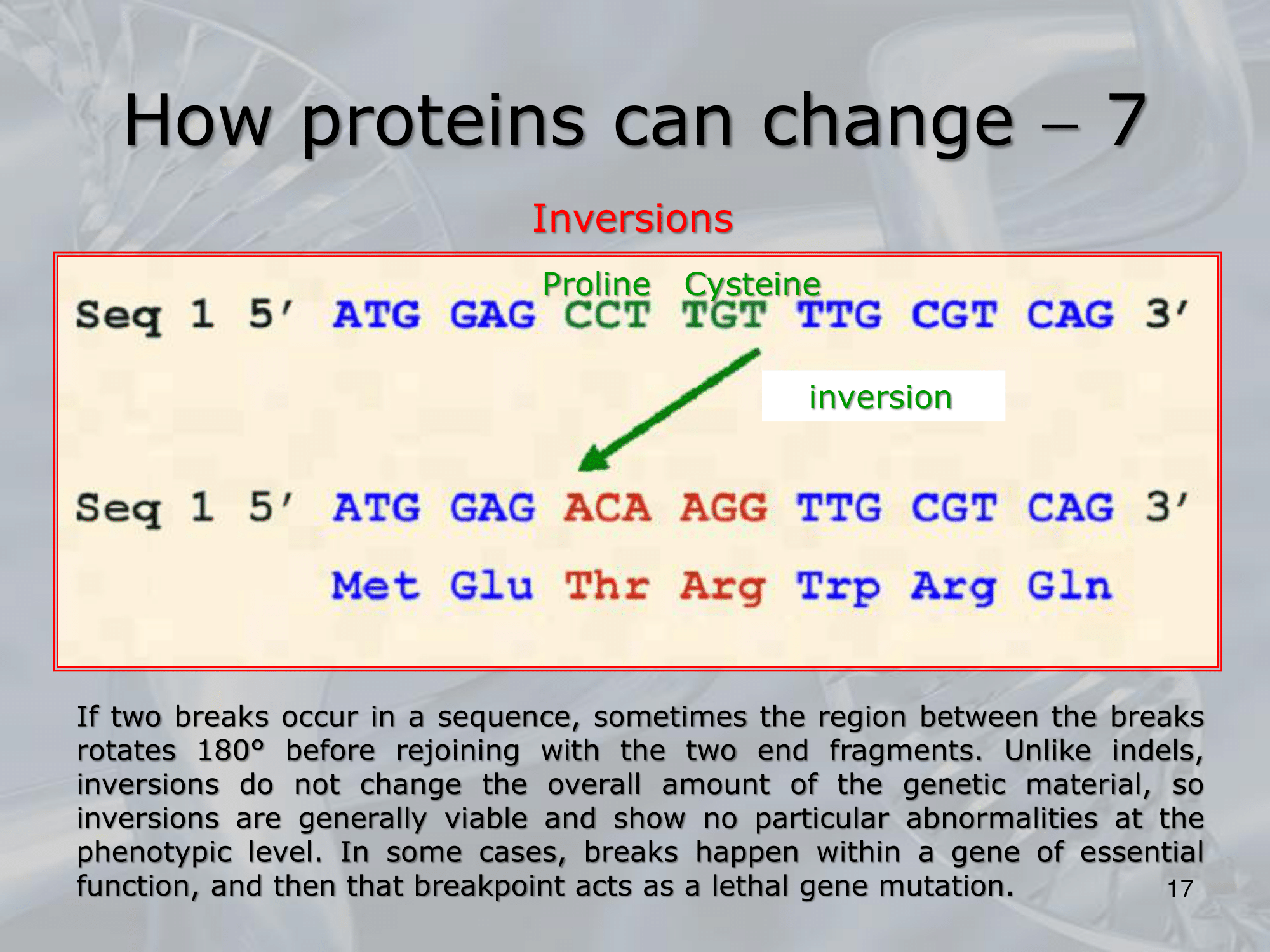
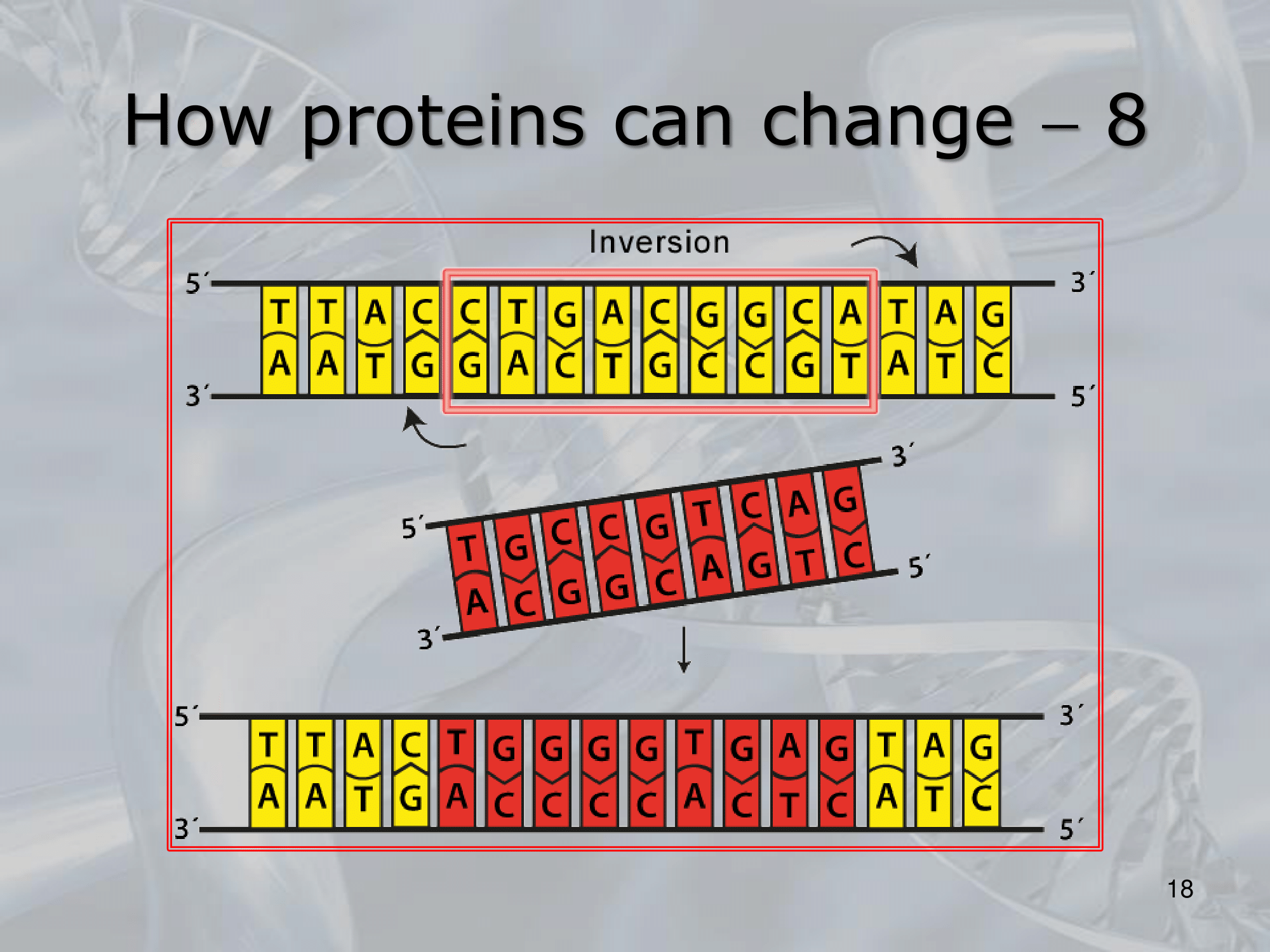
- Inversione: Un pezzetto di DNA viene invertito.
Ogniuna di queste mutazioni può avere effetti molto diversi quando si tratta di variare il codice di una catena amminoacidica.
Prendiamo come esmpio una sostituzione puntuale:
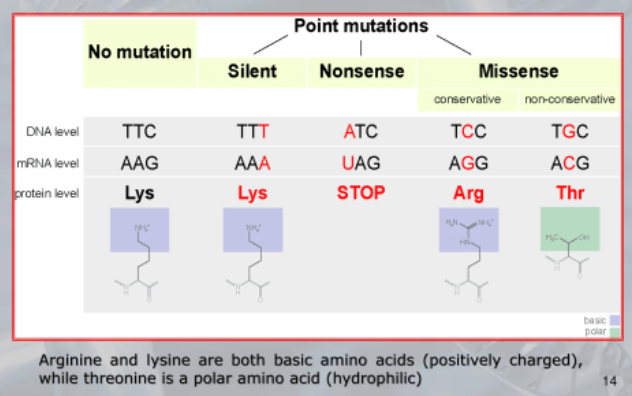
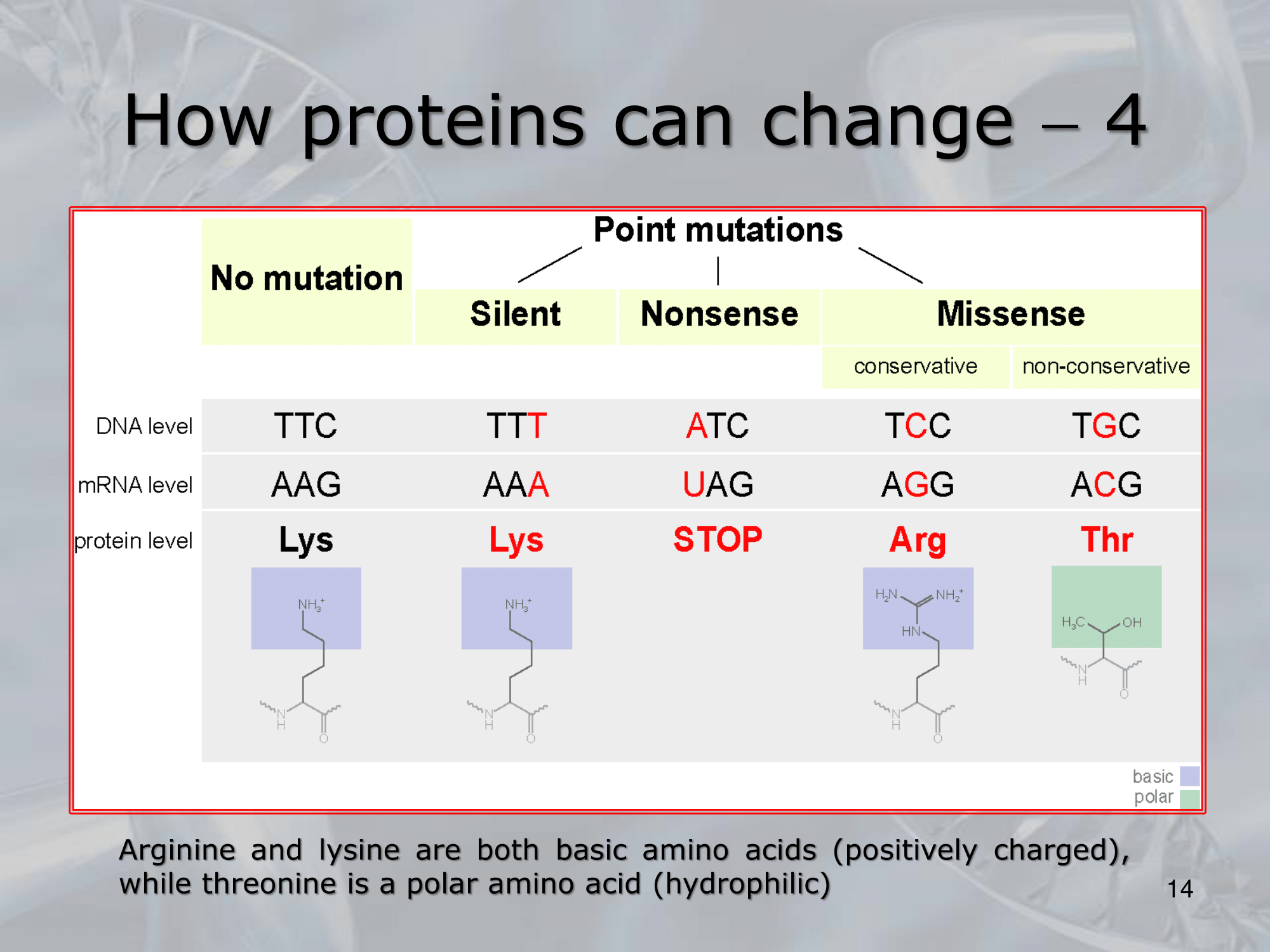
- Potremmo avere una mutazione silenziosa dove il codice di una proteina cambia, ma non cambia la sua struttura 3D.
- Una mutazione “missense conservativa”, il codice cambia variando uno o più amminoacidi ma la funzione della proteina rimane invariata.
- Una mutazione “missense non-conservativa”, la funzione della proteina varia, per esempio può capitare se cambiamo un amminoacido caricato positivamente, con un amminoacido idrofilico.
- Infine una mutazione “nosense”, per esempio se dopo una mutazione all’interno della proteina appare un codone di stop, tagliando a metà la proteina.
—————————————————————
Slides with Notes



IMPORTANTE Two Types of Point Mutation or Substitution:
- Transition: if a Purine (G,A) changes into another Purine (A,G), same with Phyrimidine
- Traslation: if a Purine (G,A) becomes a Phyrimidine (C, T), or vice-versa





IMPORTANTE Il DNA di una specie è soggetto a diverse mutazioni:
- Sostituzione Puntuale: Una singola base cambia
- Inserzione: Viene aggiunta una nuova base.
- Cancellazione: Viene tolta una base.
- Inversione: Un pezzetto di DNA viene invertito.
Ogniuna di queste mutazioni può avere effetti molto diversi quando si tratta di variare il codice di una catena amminoacidica.
Prendiamo come esmpio una sostituzione puntuale:
- Potremmo avere una mutazione silenziosa dove il codice di una proteina cambia, ma non cambia la sua struttura 3D.
- Una mutazione “missense conservativa”, il codice cambia variando uno o più amminoacidi ma la funzione della proteina rimane invariata.
- Una mutazione “missense non-conservativa”, la funzione della proteina varia, per esempio può capitare se cambiamo un amminoacido caricato positivamente, con un amminoacido idrofilico.
- Infine una mutazione “nosense”, per esempio se dopo una mutazione all’interno della proteina appare un codone di stop, tagliando a metà la proteina.