Questions
- What are Restriction Enzymes?
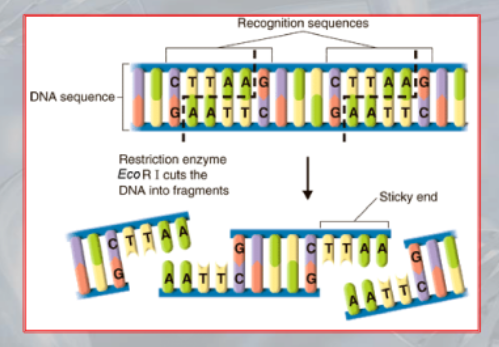
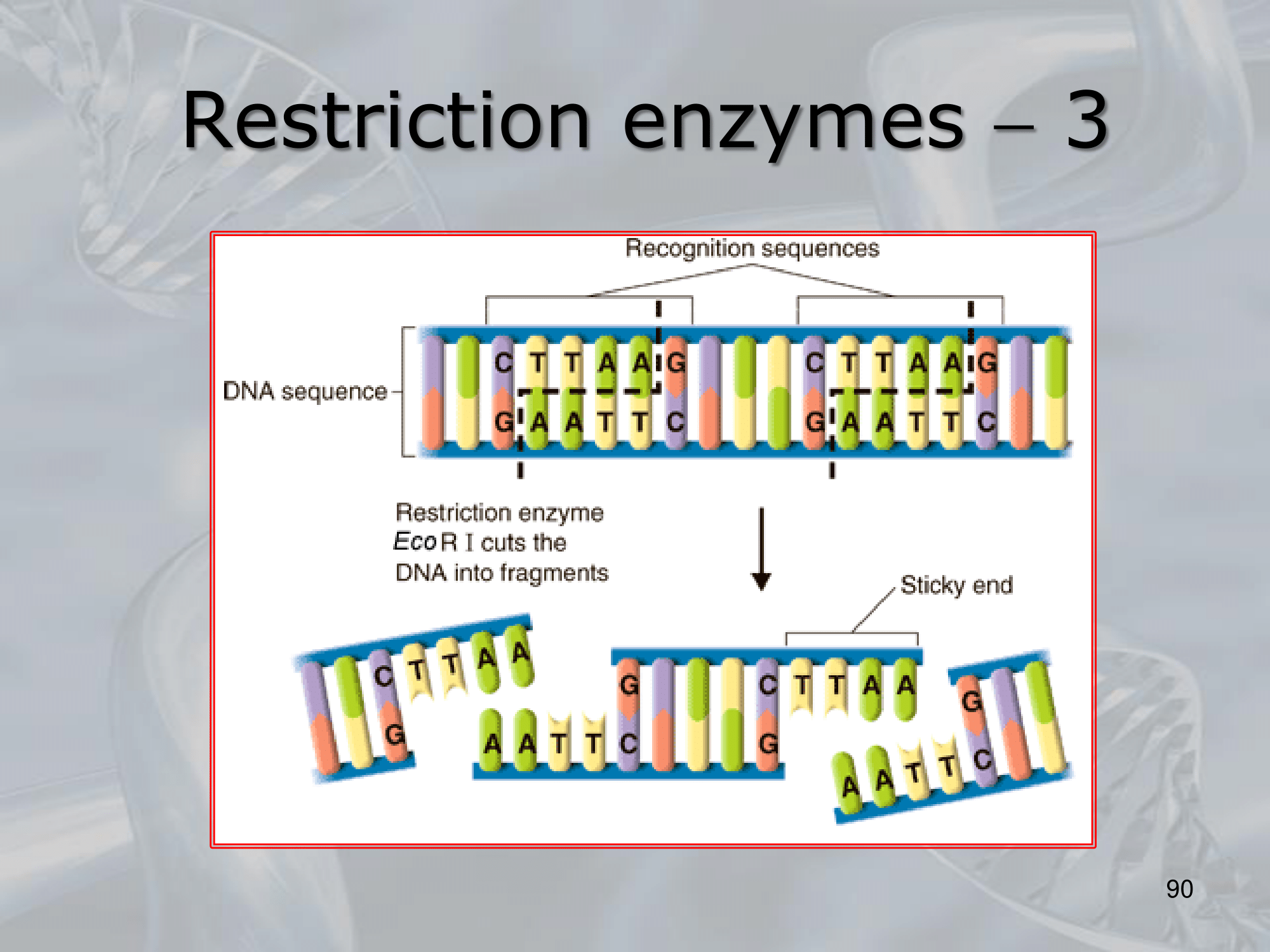
- Restriction enzymes, also known as restriction endonucleases, are enzymes that cleave DNA molecules at specific sequences, known as restriction sites. They are produced by bacteria and are part of the bacterial immune system to protect against invading foreign DNA, such as viruses.
- Restriction enzymes recognize a specific sequence of nucleotides, typically four to eight base pairs in length, and cleave the DNA at specific positions within that sequence. The resulting fragments can then be separated and analyzed using techniques such as gel electrophoresis or DNA sequencing.
- Restriction enzymes are commonly used in molecular biology for a variety of applications, such as DNA cloning, gene editing, and DNA fingerprinting. In DNA cloning, restriction enzymes are used to cut a plasmid vector and insert a desired gene fragment into the vector, creating a recombinant DNA molecule that can be transformed into a host organism. In gene editing, restriction enzymes can be used to cut a specific region of a genome, allowing for the precise deletion, insertion, or replacement of genes.
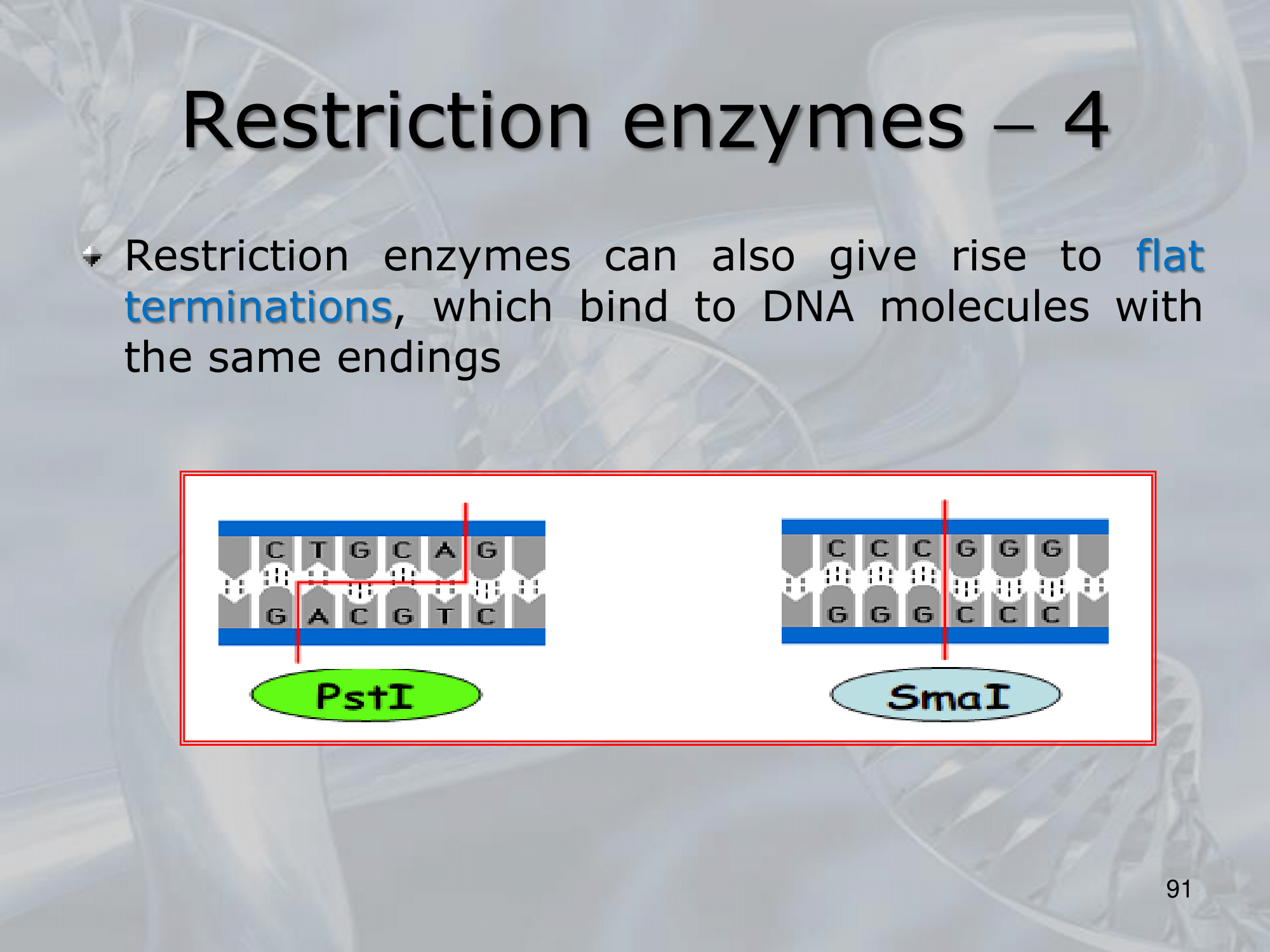
- There are many different types of restriction enzymes, each with its own specific recognition sequence and cutting site. Some restriction enzymes create blunt ends when they cut the DNA, while others create sticky ends with overhanging nucleotides that can anneal to complementary sequences, allowing for the creation of recombinant DNA molecules.
- What is a Restriction Map?
- A restriction map is a graphical representation of the locations of restriction sites along a DNA molecule.
It shows the sizes and relative positions of the fragments generated by cleavage with one or more restriction enzymes. - A restriction map is created by digesting a DNA sample with one or more restriction enzymes, separating the resulting fragments by size using gel electrophoresis, and then analyzing the pattern of fragment sizes.
The sizes of the fragments are determined by comparing them to a set of known DNA standards of known sizes, which are also separated by gel electrophoresis. - ==Restriction maps can be used to identify unknown DNA samples, confirm the identity and purity of DNA samples, and map the location of genes and other functional elements within a DNA molecule.
They can also be used to compare the DNA sequences of different individuals or species, as differences in restriction site patterns can indicate variations in the underlying DNA sequence==. - Overall, restriction maps are an important tool in molecular biology and genetics, and have contributed greatly to our understanding of DNA structure, function, and evolution.
- A restriction map is a graphical representation of the locations of restriction sites along a DNA molecule.
IMPORTANTE
IMPORTANTE Restriction Enzymes Enzymes that can be used as “scissors”, which allow biologists to cut and paste the DNA molecules. The “sticky ends“ leaved by the enzyme can be rejoined by another enzyme called ligase.
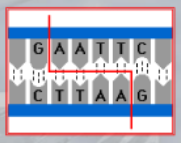
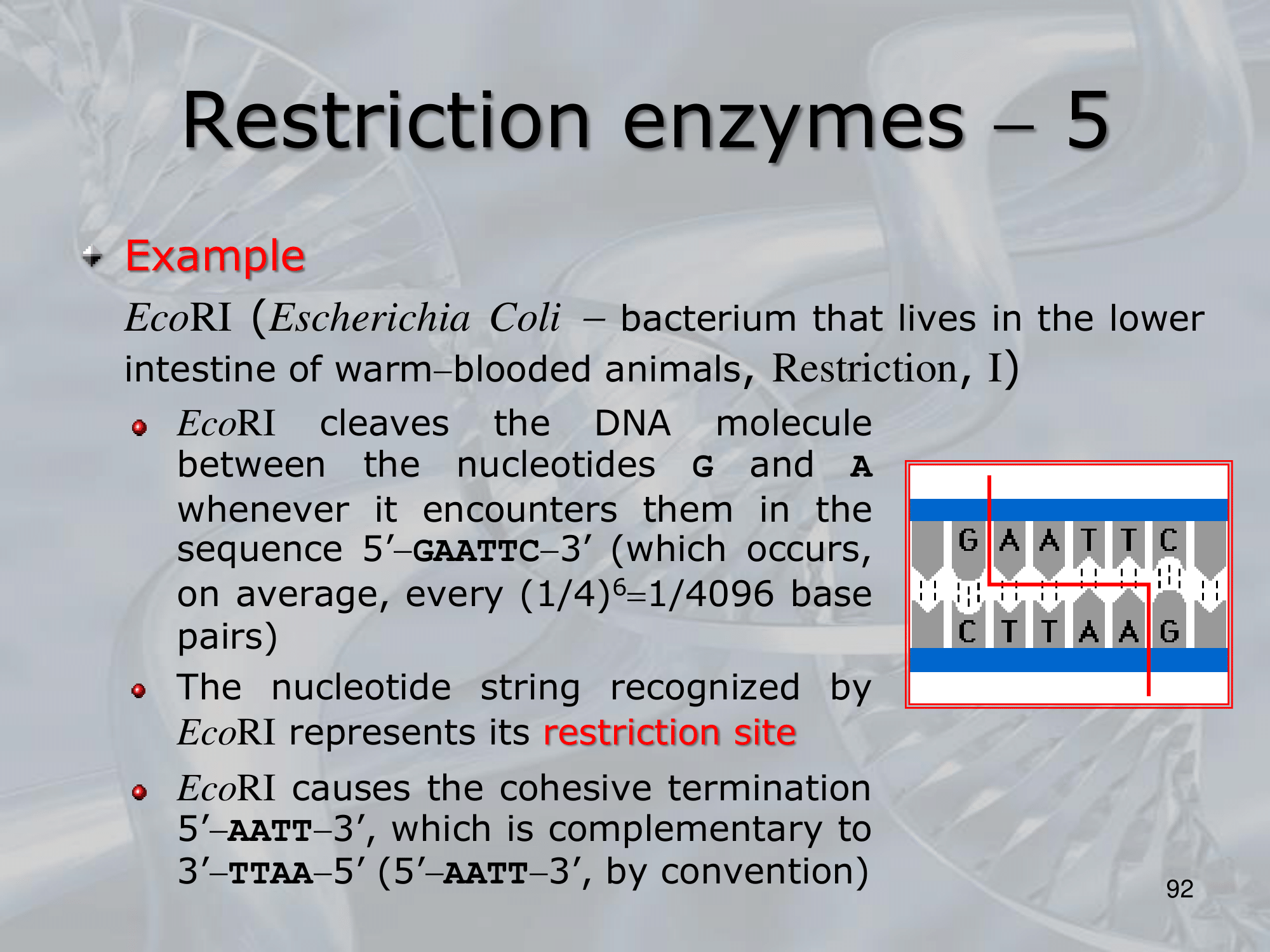
IMPORTANTE Restriction Site The site recognized by the restriction enzyme to be cut. For example, the EcoRI cuts when it finds the sequence GAATTC, which is its restriction site (it leaves the AATT as the stiky-end).
IMPORTANTE Restriction Maps A map showing the position of restriction cuts, within a particular DNA sequence
Slides with Notes


IMPORTANTE Restriction Enzymes Enzymes that can be used as “scissors”, which allow biologists to cut and paste the DNA molecules. The “sticky ends“ leaved by the enzyme can be rejoined by another enzyme called ligase.
IMPORTANTE Restriction Site The site recognized by the restriction enzyme to be cut. For example, the EcoRI cuts when it finds the sequence GAATTC, which is its restriction site (it leaves the AATT as the stiky-end).
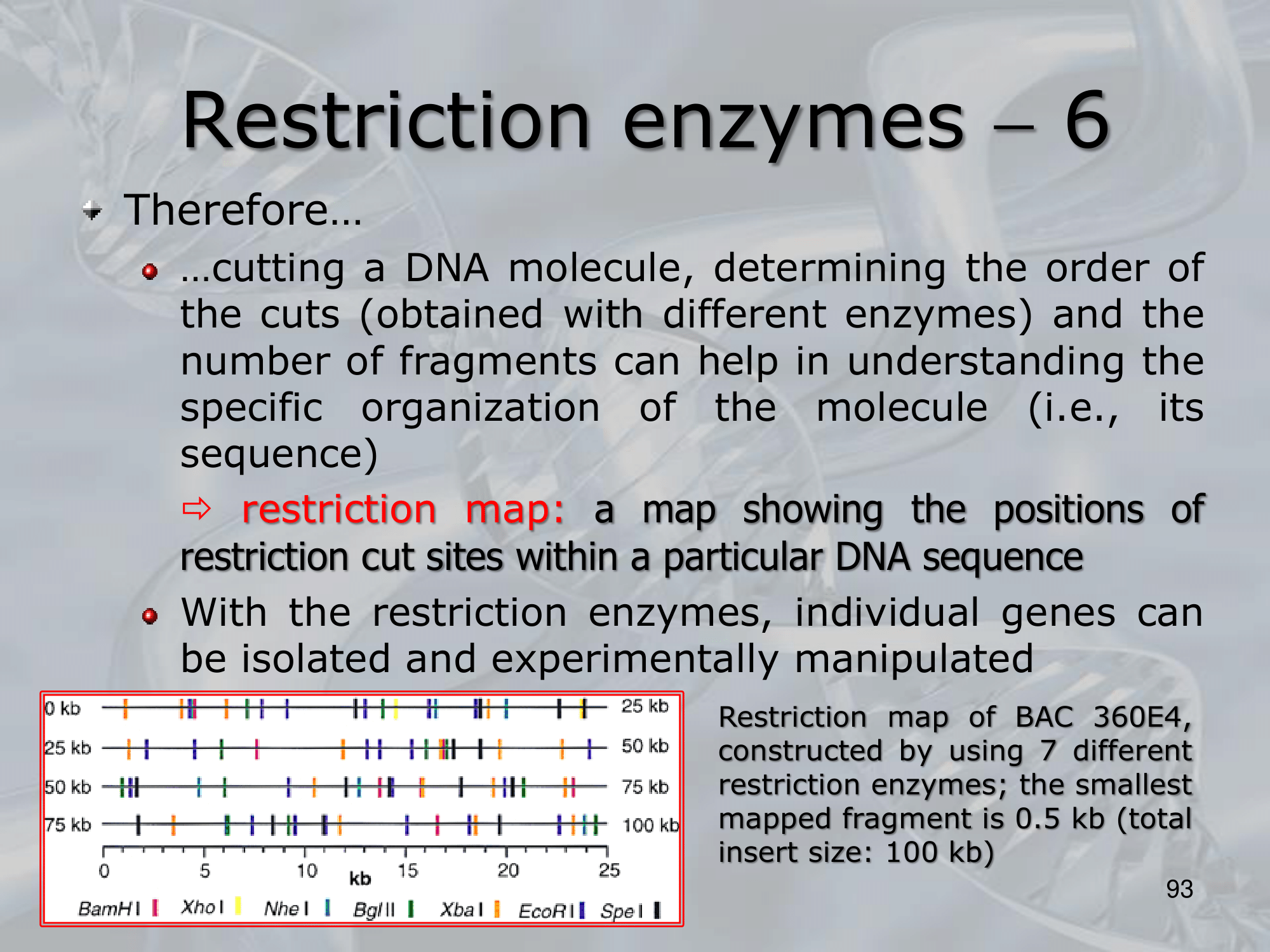


IMPORTANTE Restriction Maps A map showing the position of restriction cuts, within a particular DNA sequence