Questions
- What is Blotting and Hybridization?
- Blotting and hybridization are techniques used to detect and analyze specific DNA or RNA sequences within a complex mixture of nucleic acids.
- Blotting involves transferring nucleic acid fragments from a gel matrix onto a solid membrane, such as nitrocellulose or nylon, by capillary action or electroblotting.
This creates a replica of the gel matrix on the membrane, with the nucleic acid fragments immobilized on the surface of the membrane. - ==Hybridization involves using a labeled nucleic acid probe that is complementary to a specific target sequence within the immobilized nucleic acid fragments on the membrane.
The probe is typically labeled with a radioactive or fluorescent tag that allows for detection and visualization of the hybridization signal==. - When the labeled probe is added to the membrane, it will hybridize with any immobilized nucleic acid fragments that contain a complementary sequence. The membrane is then washed to remove any unbound probe, and the labeled probe that has hybridized to the target sequence is detected by autoradiography or fluorescence.
- Blotting and hybridization are commonly used in molecular biology and genetics for a variety of applications, such as DNA and RNA sequencing, gene expression analysis, and detection of mutations and genetic variations.
These techniques allow for the detection and analysis of specific nucleic acid sequences within complex mixtures, providing valuable information about the structure and function of genes and other genetic elements.
IMPORTANTE
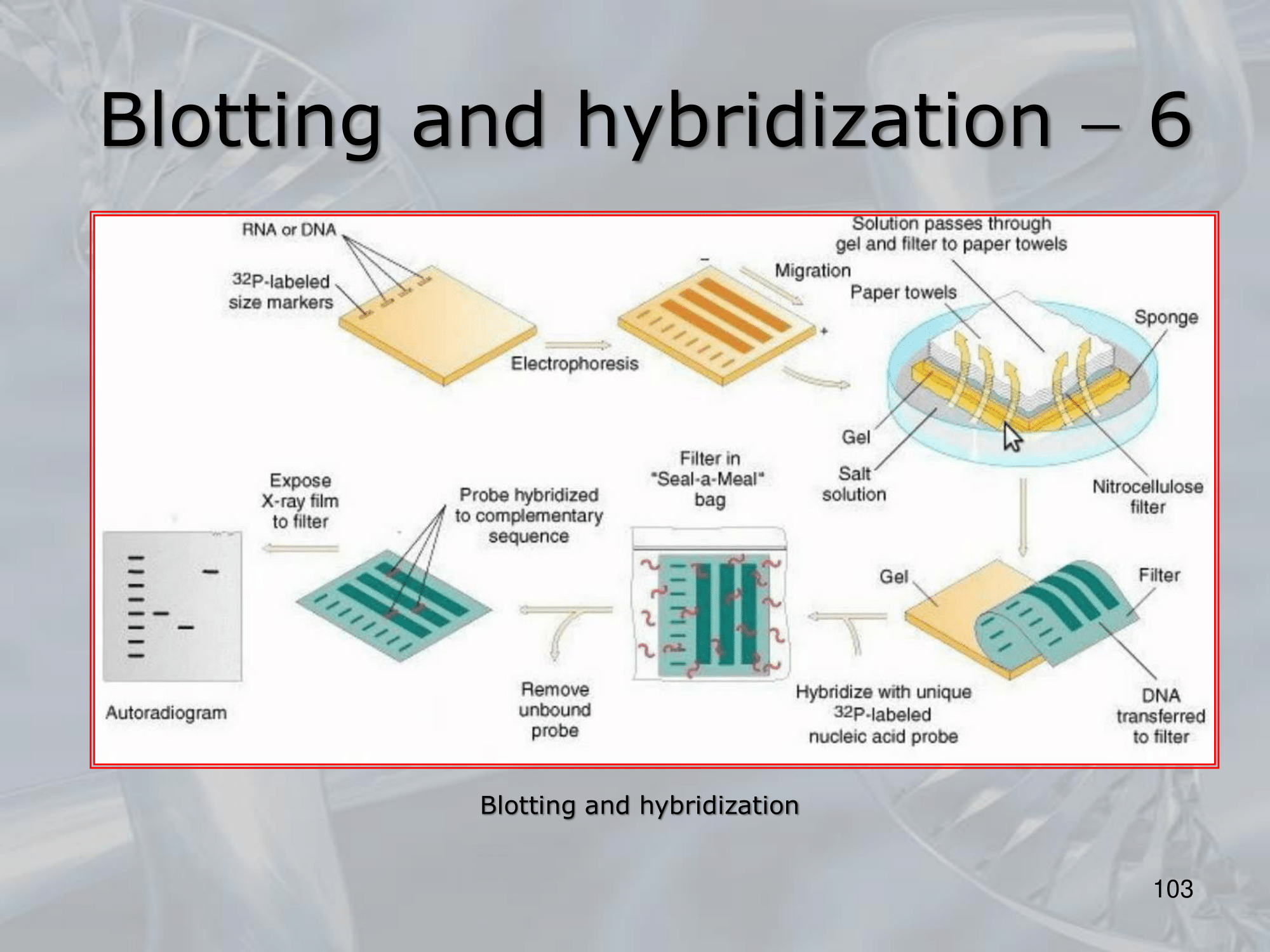
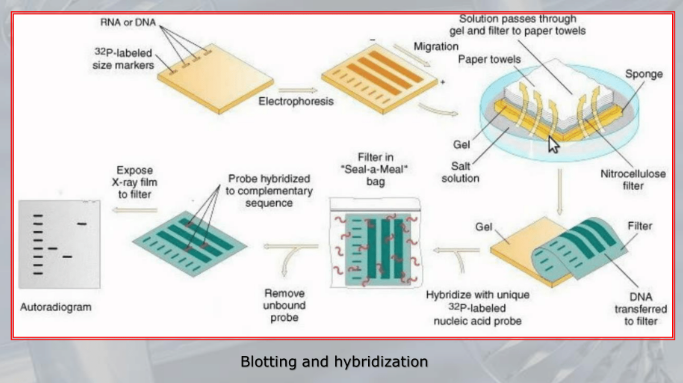
IMPORTANTE Blotting and Hybridization Process used to search if a particular gene (or more general, a particular DNA fragment) is present in the DNA fragments we are studying
- Blotting phase: Polynucleotides (DNA or RNA) are transferred from the gel to a solid support, which also produces DNA denaturation, the DNA moleucules remain attached to the membrane in the position previously occupied by the gel.
- Cooking Proecess: Fixes rthe DNA fragments to the membrane.
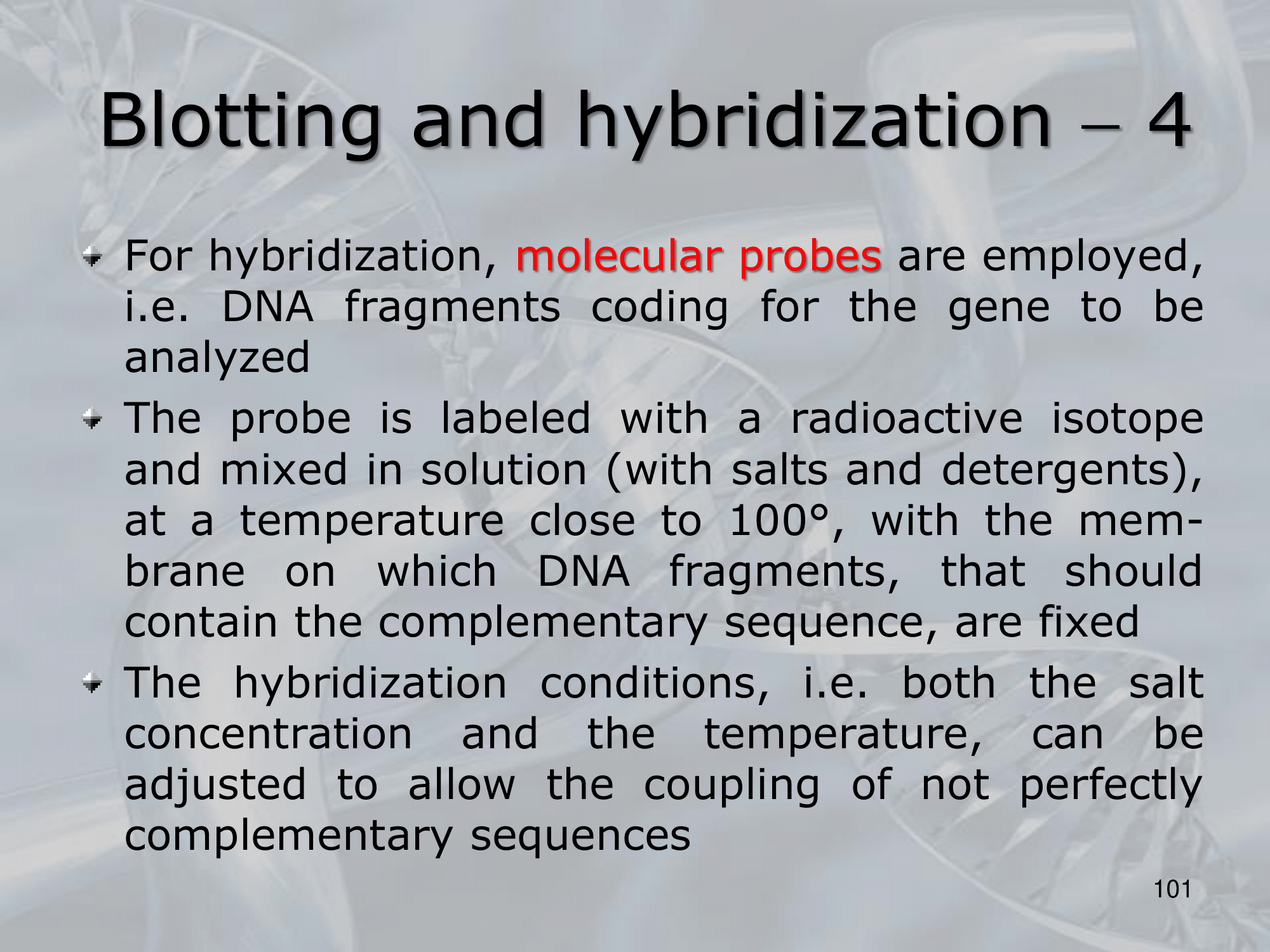
- Hybridization: We label DNA fragments that code for known genes, with a radioactive istope, creating what we call molecular probes, then we mix this probes in a solution at 100° C, with the blotted membrane, this will fix the probes to the complementary DNA on the blotted membrane.
- We wash away the molecular probes that did not bind (in the blotting memebrane there were no cDNA of this probes), and we verify which genes are present on the original DNA on the blotted membrane, we can see them because we nade them radiocative.
NOTE: The hybridization condition: salt concentration and temperature, can be changed to allow the coupling of not perfect complementary sequences. ~Ex.: changing the hybridization conditions we can allow the two fragments ATTCG and TATGC to bind, even if they are not perfect complementary sequences
Slides with Notes
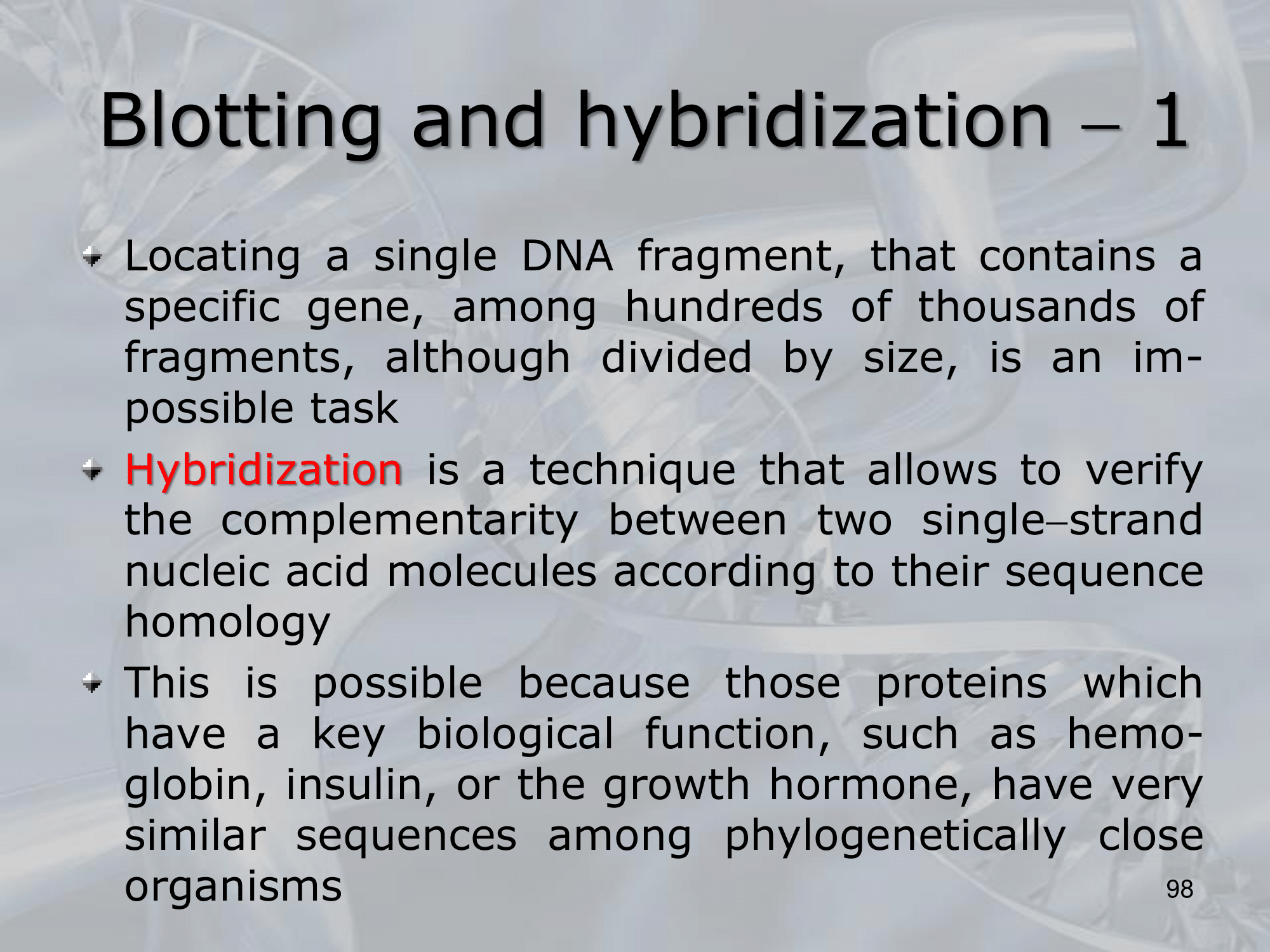
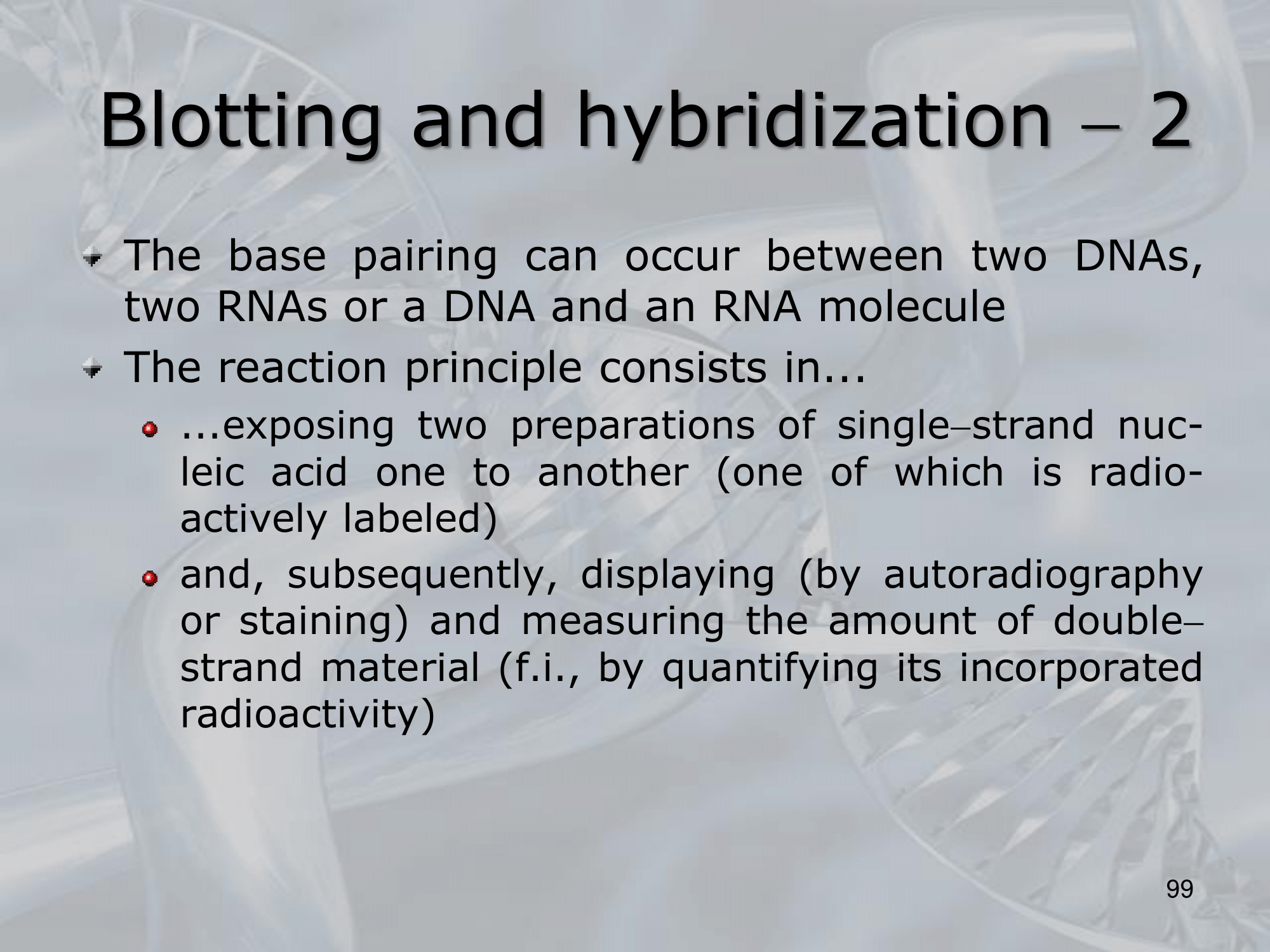
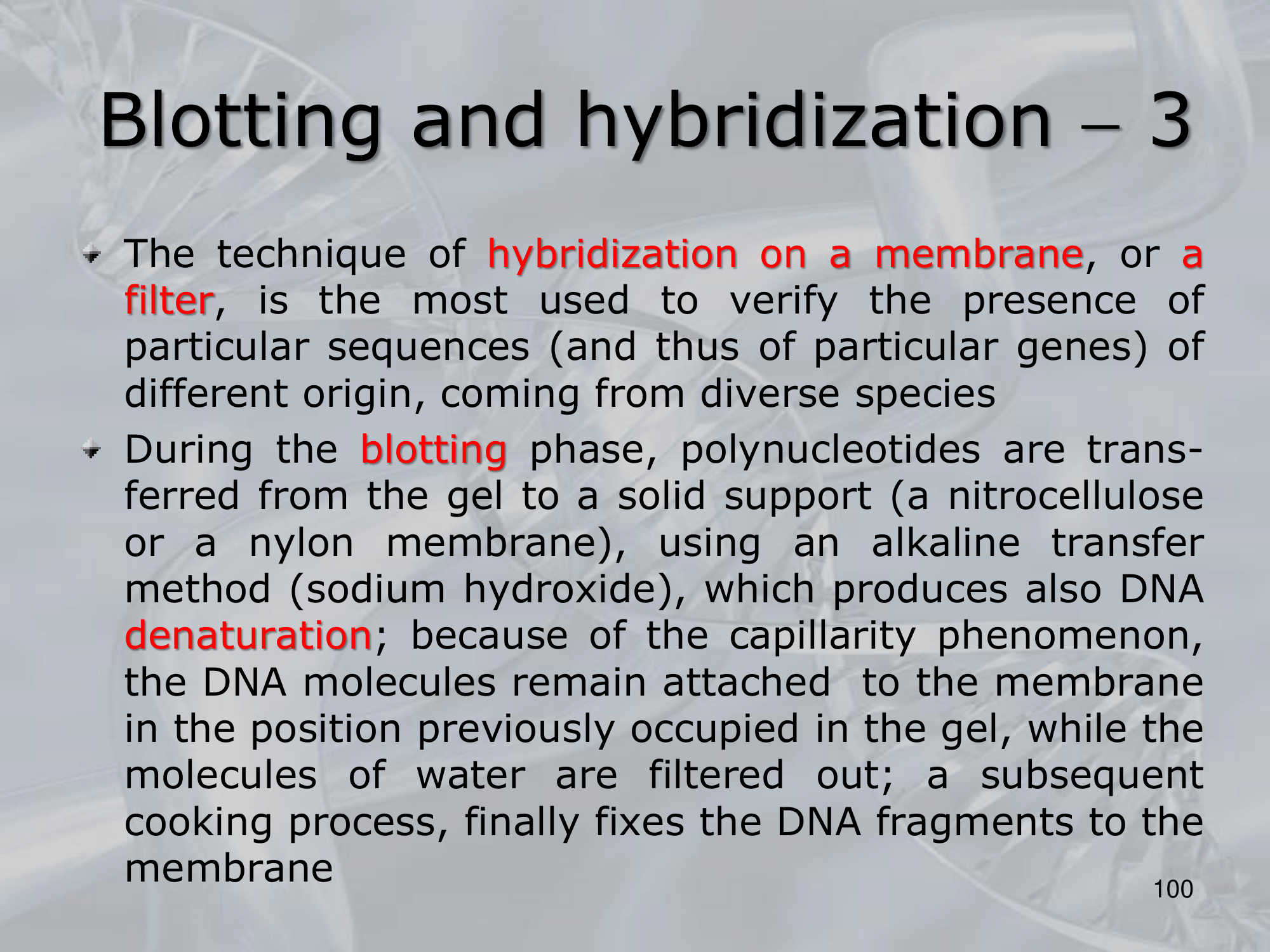



IMPORTANTE Blotting and Hybridization Process used to search if a particular gene (or more general, a particular DNA fragment) is present in the DNA fragments we are studying
- Blotting phase: Polynucleotides (DNA or RNA) are transferred from the gel to a solid support, which also produces DNA denaturation, the DNA moleucules remain attached to the membrane in the position previously occupied by the gel.
- Cooking Proecess: Fixes rthe DNA fragments to the membrane.
- Hybridization: We label DNA fragments that code for known genes, with a radioactive istope, creating what we call molecular probes, then we mix this probes in a solution at 100° C, with the blotted membrane, this will fix the probes to the complementary DNA on the blotted membrane.
- We wash away the molecular probes that did not bind (in the blotting memebrane there were no cDNA of this probes), and we verify which genes are present on the original DNA on the blotted membrane, we can see them because we nade them radiocative.
NOTE: The hybridization condition: salt concentration and temperature, can be changed to allow the coupling of not perfect complementary sequences. ~Ex.: changing the hybridization conditions we can allow the two fragments ATTCG and TATGC to bind, even if they are not perfect complementary sequences