Questions
- How can we use Neural Networks to Predict the Secondary Strucutre of a Protien?
- Neural networks can be used to predict the secondary structure of a protein from its amino acid sequence.
Here is a general overview of the process:- Data preparation: A dataset of known protein structures is needed for training the neural network.
The dataset should include protein sequences and their corresponding secondary structure annotations (e.g., alpha helix, beta sheet, or coil). - Feature extraction: The amino acid sequence of each protein in the dataset is converted into a set of numerical features that can be used as input to the neural network.
Common features include the identity of each amino acid, its position in the sequence, and physicochemical properties such as hydrophobicity and charge. - Model architecture: A neural network architecture is designed and trained on the dataset.
The architecture typically consists of multiple layers of nodes (neurons), each of which performs a nonlinear transformation on the input features.
The final layer of the network produces an output vector that represents the predicted probability of each type of secondary structure element at each position in the protein sequence. - Training: The neural network is trained on the dataset using a supervised learning approach.
During training, the network learns to minimize the difference between its predicted secondary structure and the true secondary structure annotations in the dataset.
This is typically done using a loss function such as cross-entropy or mean squared error. - Evaluation: Once the network is trained, it can be used to predict the secondary structure of new proteins.
The accuracy of the predictions is typically evaluated using metrics such as sensitivity, specificity, and F1 score, which measure the proportion of correctly predicted secondary structure elements.
- Data preparation: A dataset of known protein structures is needed for training the neural network.
- Neural networks have been shown to be highly effective for protein secondary structure prediction, and have achieved state-of-the-art performance on benchmark datasets.
However, they require a large amount of training data and computational resources, and may be prone to overfitting if the dataset is too small or too biased.
- Neural networks can be used to predict the secondary structure of a protein from its amino acid sequence.
—————————————————————
IMPORTANTE
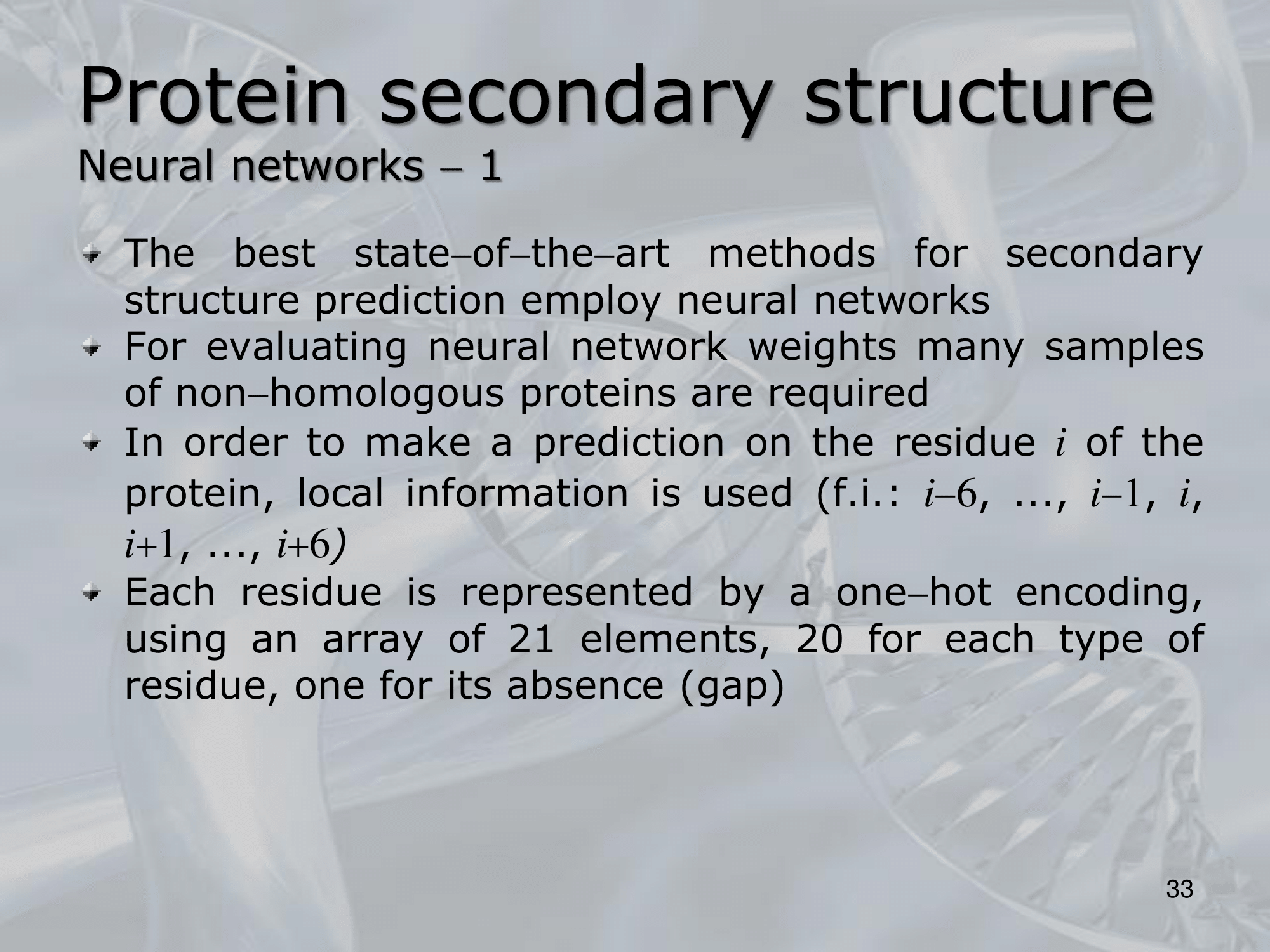
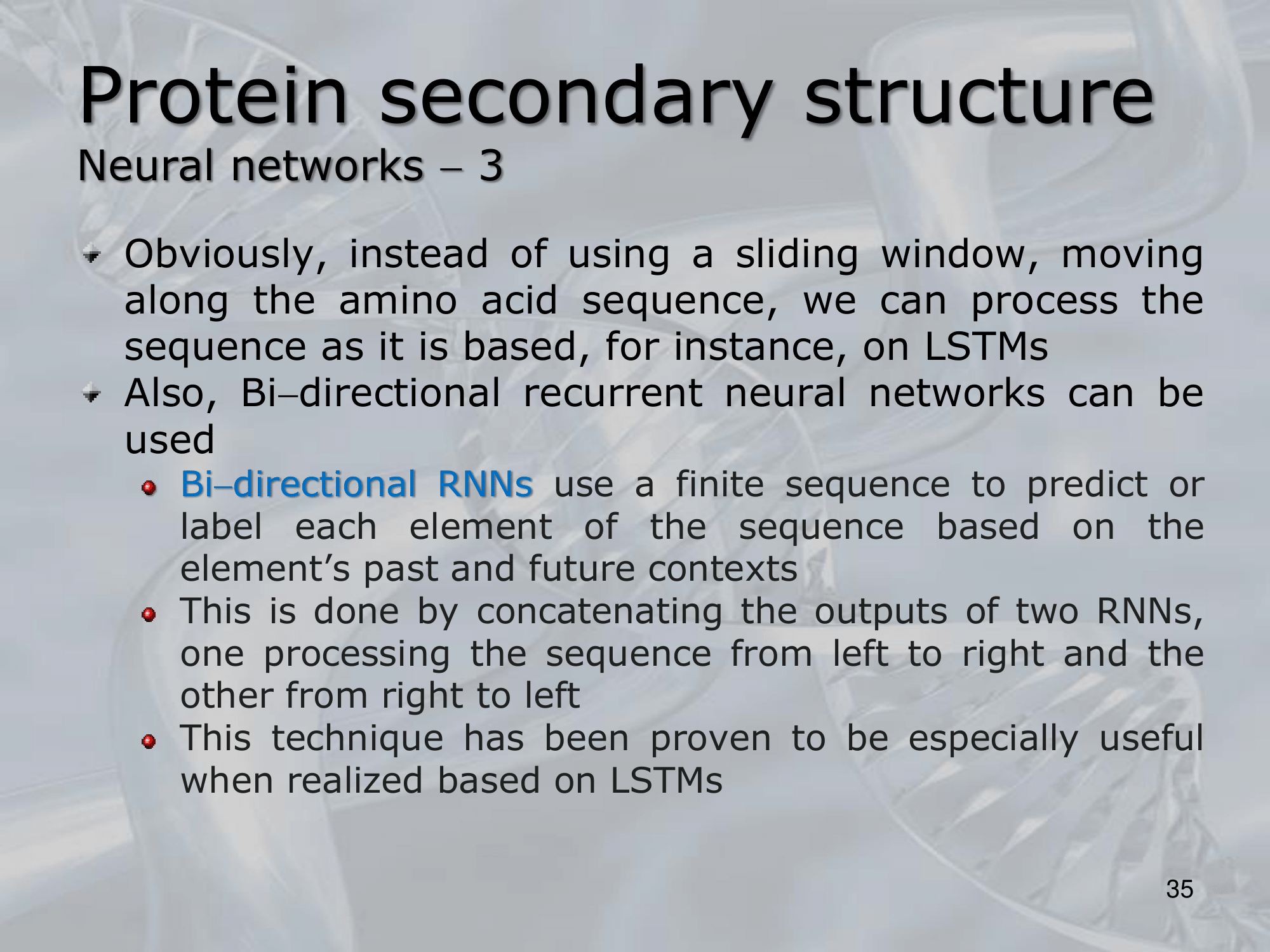
IMPORTANTE Neural Networks:
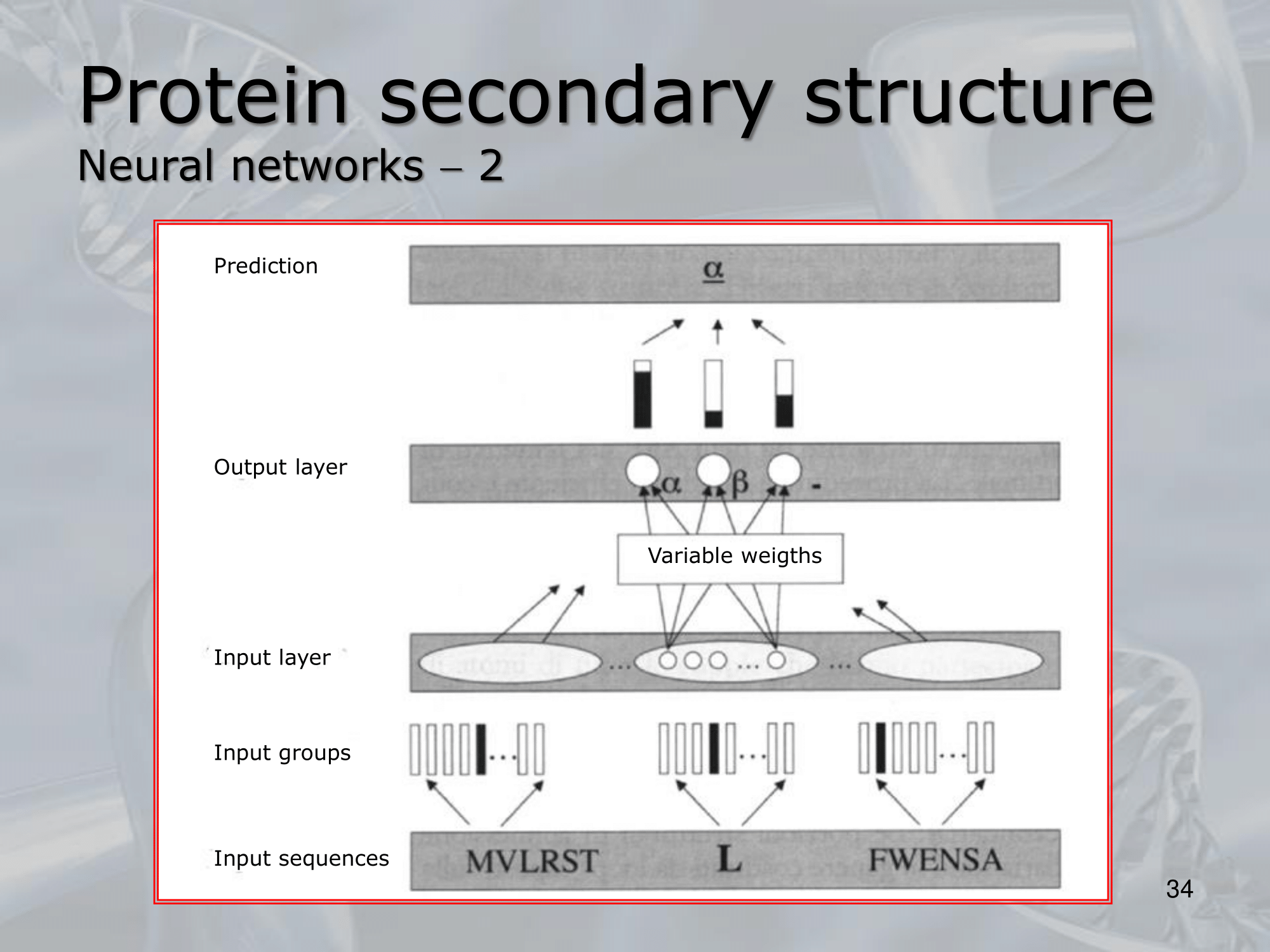
Like the GOR method we can take a sliding window and evaluate the central residue. Or we can use an LSTM NN (Long Shor Term Memory Neural Network) and pass the whole protein code, as output it will still give use a vector of same length with the probabilities of each residue of being one of the four “classes”: a helic, a sheet, a turn or a random coil.
—————————————————————
Slides with Notes