Questions
- What are Isochores?
- ==Isochores are long stretches of DNA in a genome that have a relatively uniform base composition.
Specifically, they are regions of DNA that have a high GC content or a low GC content, with a sharp transition between the two regions==.
These regions can be several hundred kilobases long and can account for a significant portion of the genome. - Isochores were first identified in the 1980s, and their study has provided important insights into genome organization and evolution.
It is now recognized that genomes of most eukaryotes are partitioned into isochores, and that there are characteristic differences in the GC content and gene density of isochores. - Isochores with a high GC content tend to be gene-rich and are often associated with important functional regions of the genome, such as gene clusters and regulatory elements.
On the other hand, isochores with a low GC content tend to be gene-poor and are often associated with repetitive DNA sequences and other non-coding regions of the genome. - The study of isochores has also shed light on the evolutionary history of genomes.
It is now thought that the ancestral eukaryotic genome had a relatively uniform GC content, and that the partitioning of the genome into isochores occurred as a result of a combination of mutation, recombination, and selective pressures.
In particular, it is thought that isochores with a high GC content may have evolved as a result of the selective pressure for efficient and accurate transcription, while isochores with a low GC content may have evolved as a result of selective pressure for genome stability. - Overall, the study of isochores has provided important insights into genome organization and evolution, and continues to be an active area of research in the field of genomics.
- ==Isochores are long stretches of DNA in a genome that have a relatively uniform base composition.
—————————————————————
IMPORTANTE
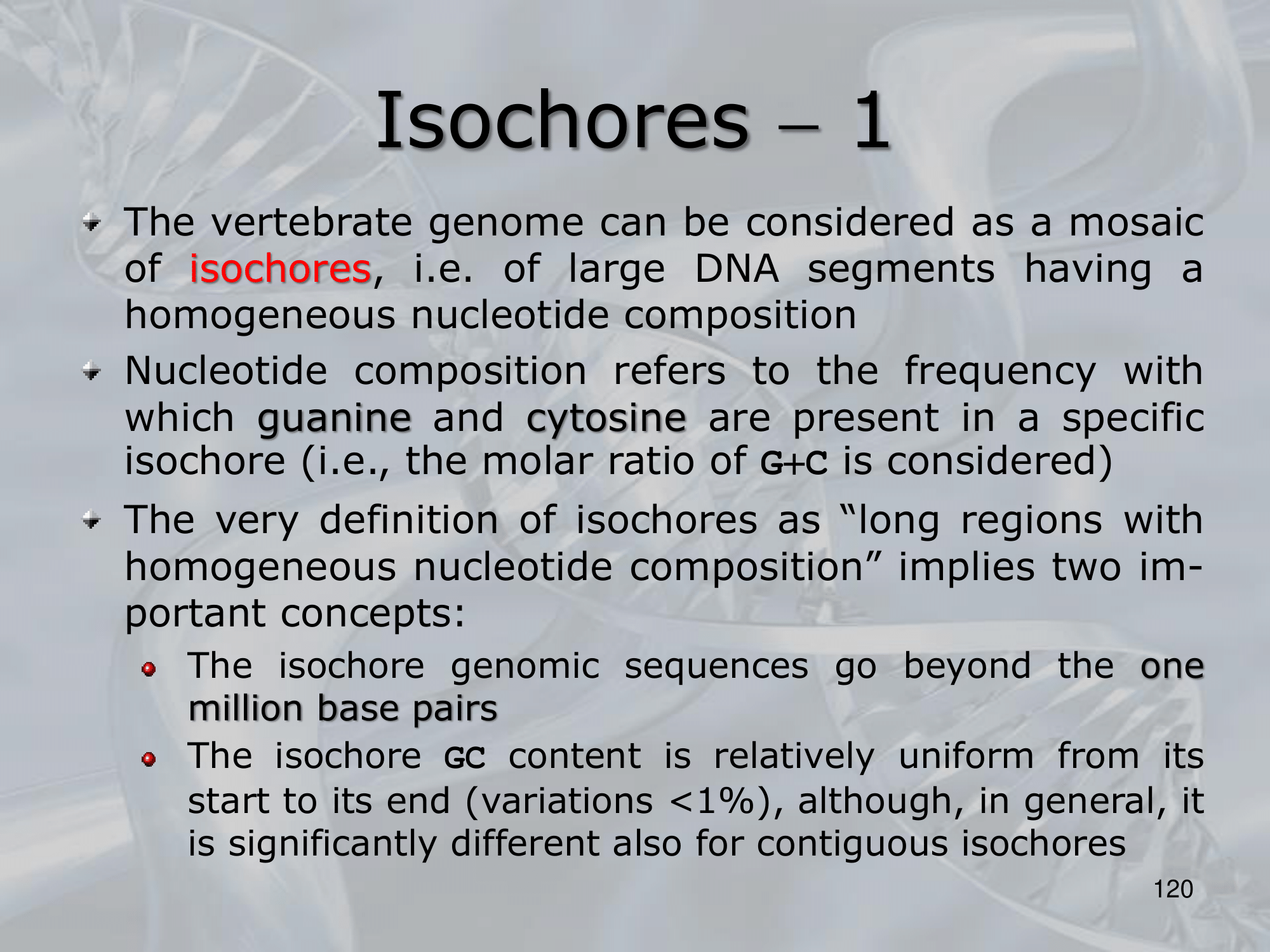
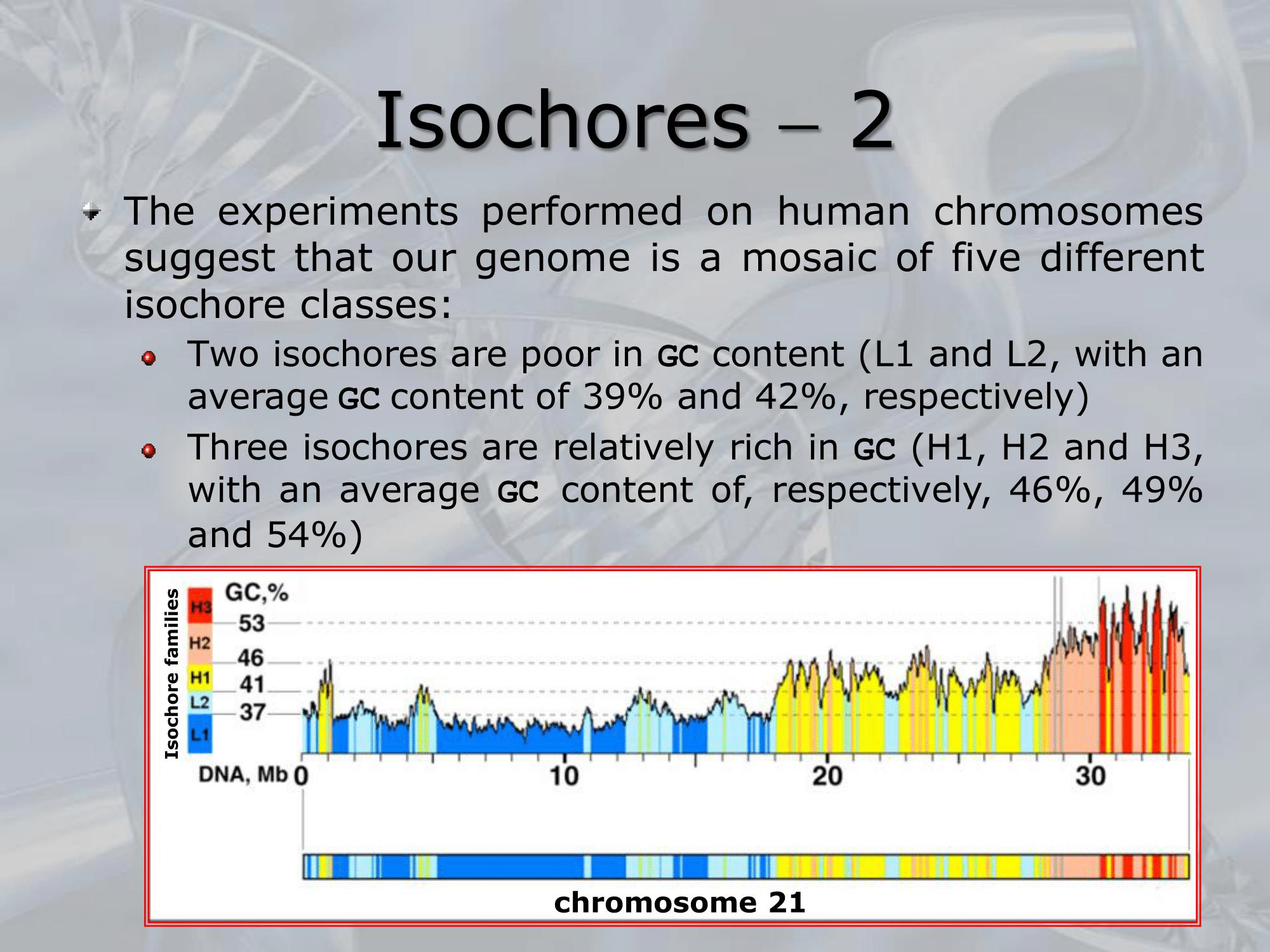
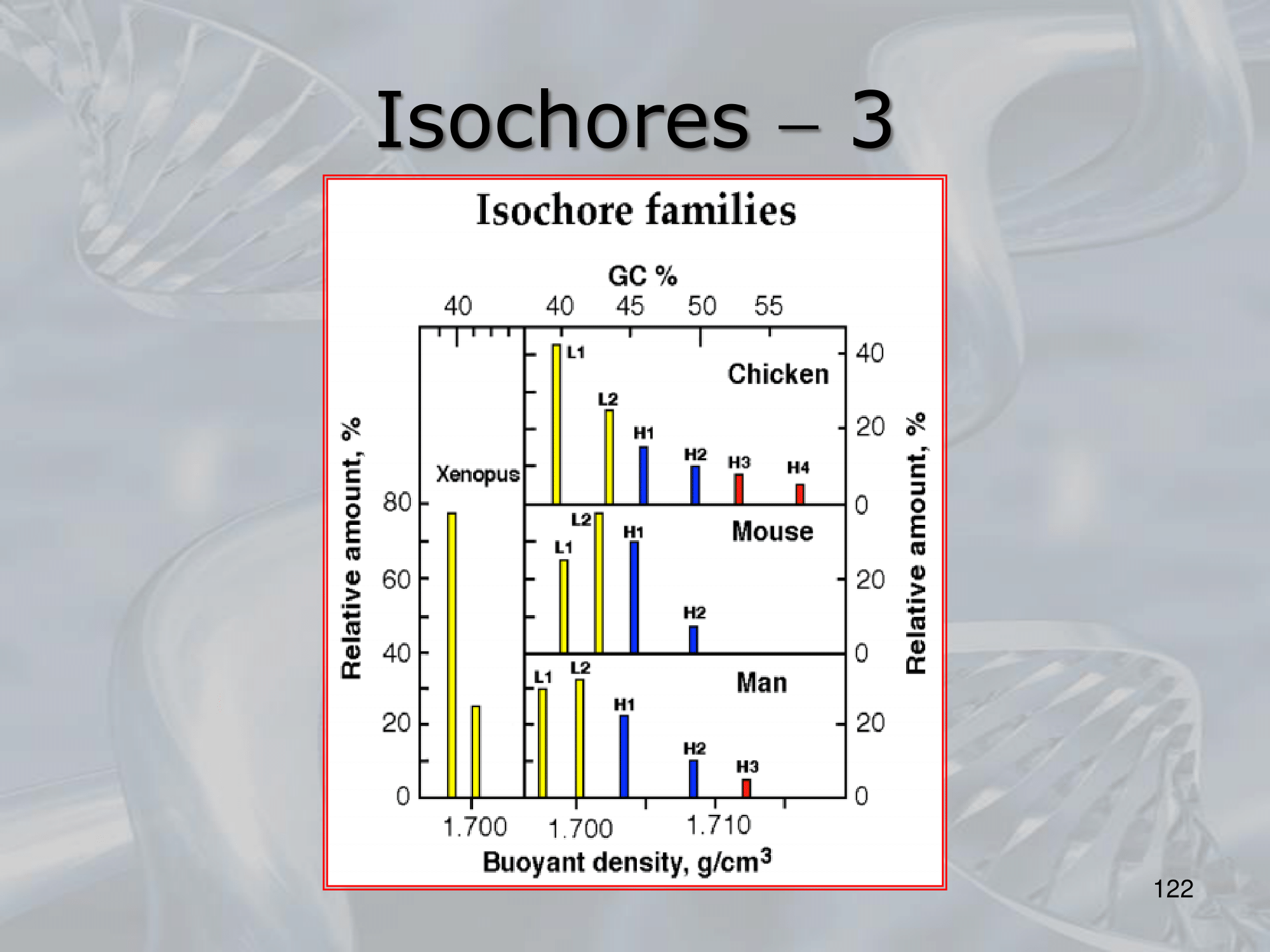
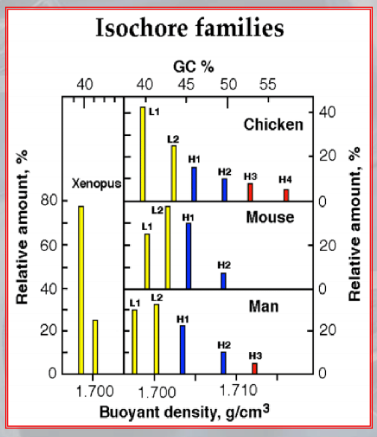
IMPORTANTE In the vertebrate genome we can find 5 types of isochores. Each isochore is a portion of DNA (above one million base pairs) characterized by a specific percentage of avarege GC Content.
- L1 isochore: average of of GC content (poor in GC content)
- L2 isochore: average of of GC content (poor in GC content)
- H1 isochore: average of of GC content (high in GC content)
- H2 isochore: average of of GC content (high in GC content)
- H3 isochore: average of of GC content (high in GC content)
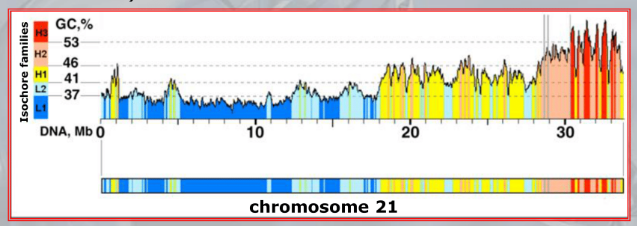
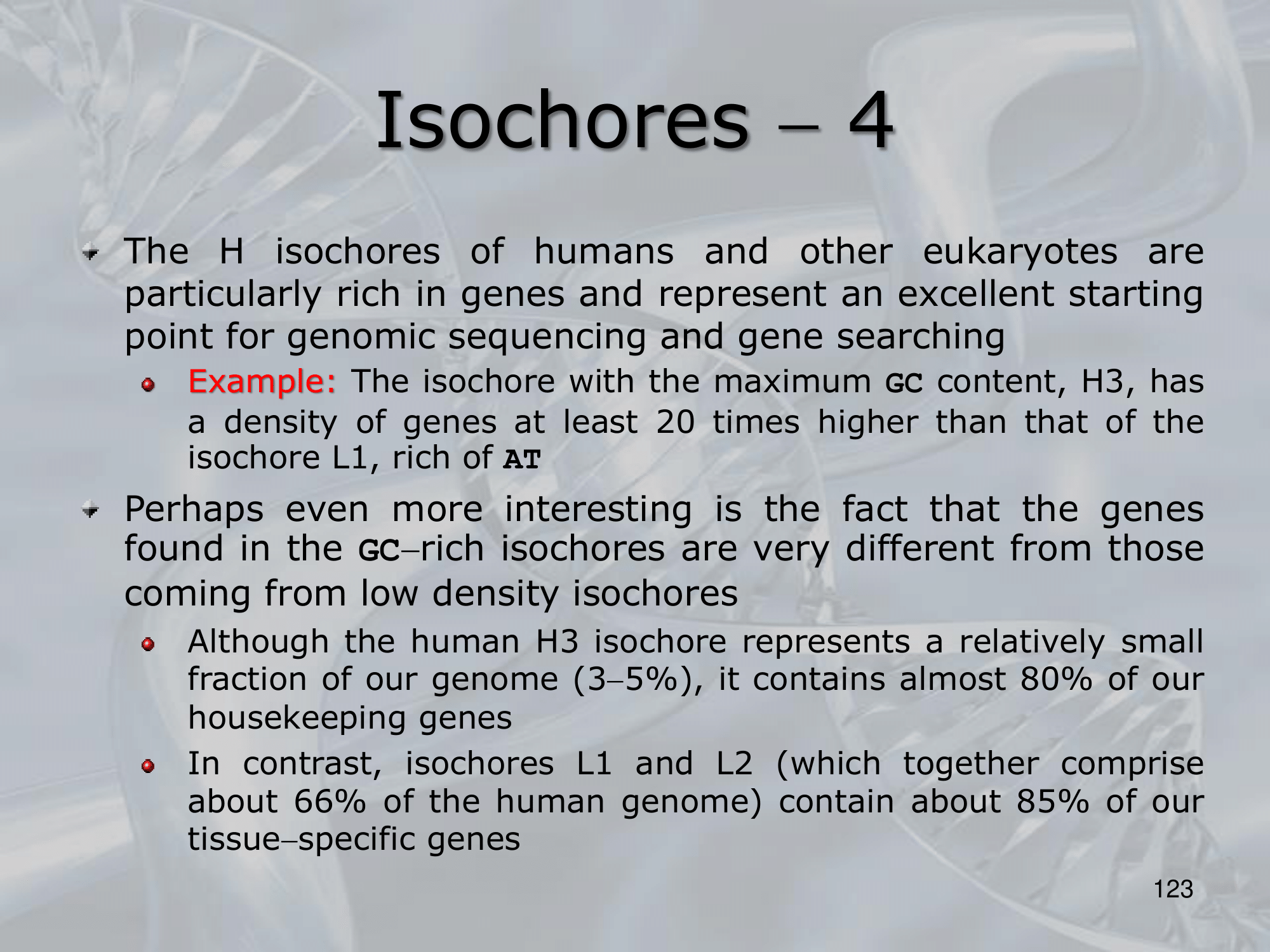
Each isochore is also characterized by a different percentage of gene density. ~Ex.: the H3 isochore, rich of GC () has a density of genes at least 20 times higher than the L1 ischore, which is rich of AT.
Also it is intresting to note that the type of genes are very different in different isochores:
- H3 contains of our housekeeping genes
- L1 and L2 contain together of our tissue-specific genes
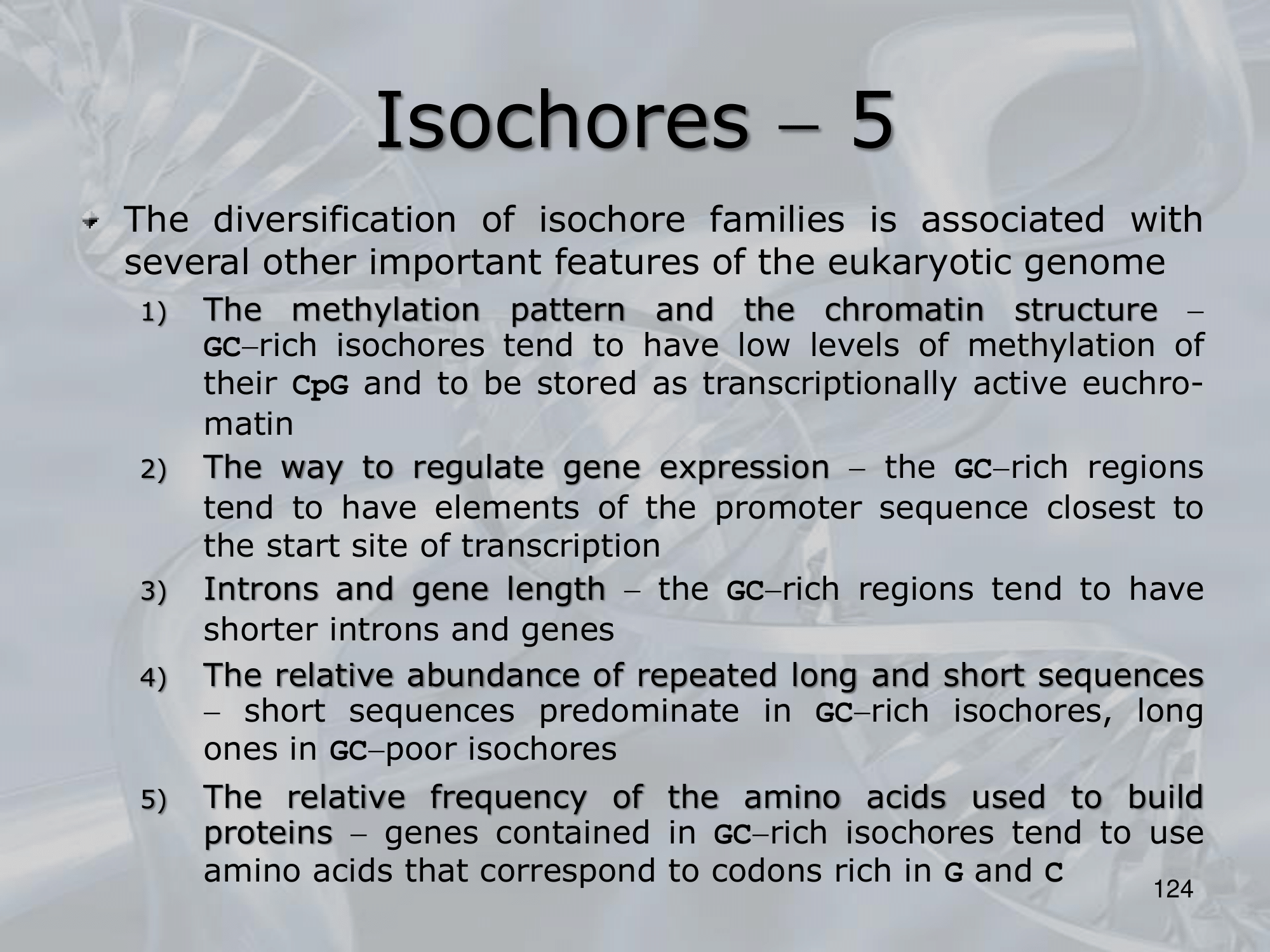
Also to note that isochore with higher GC content (H1, H2, H3) have:
- Smaller-sized repeated sequences
- Lower level of methylation, due the high presence of CpG islands.
- Intros and exons are much smaller in size.
- Promoters are closer to the relative genes.
While lower GC content isochores (L1, L2) have:
- repeated sequences which are on average much longer.
—————————————————————
Slides with Notes