Questions
- What are Synonimous and Non-Synonymous Substitutions, what are their Differences?
- ==Synonymous substitutions, also known as silent substitutions, are nucleotide changes in the DNA sequence that do not result in an amino acid change in the corresponding protein==.
This is because the genetic code is degenerate, meaning that multiple codons can code for the same amino acid.
Synonymous substitutions do not affect the protein sequence or function and are often assumed to be selectively neutral. - Non-synonymous substitutions, on the other hand, are nucleotide changes in the DNA sequence that do result in an amino acid change in the corresponding protein.
Non-synonymous substitutions can be further classified into two types: missense and nonsense substitutions.- ==Missense substitutions result in the replacement of one amino acid with another==.
- ==Nonsense substitutions result in the premature truncation of the protein due to the creation of a stop codon==.
- The primary difference between synonymous and non-synonymous substitutions is that synonymous substitutions do not result in an amino acid change and are often assumed to be selectively neutral, while non-synonymous substitutions can have a range of effects on protein function, including changes in activity, stability, folding, and interactions.
Non-synonymous substitutions are more likely to be subject to natural selection, as they can affect the fitness of an organism.
- ==Synonymous substitutions, also known as silent substitutions, are nucleotide changes in the DNA sequence that do not result in an amino acid change in the corresponding protein==.
- What are the Categories of Degenerate Sites within a Codon?
- Within a codon, there are three nucleotide positions.
The first and second positions are usually more conserved and undergo fewer substitutions, while the third position is more degenerate and undergoes more substitutions without affecting the amino acid sequence of the protein.
The degeneracy of the third position is due to the presence of multiple codons that encode the same amino acid, which are referred to as synonymous codons. - The categories of degenerate sites within a codon are:
- ==Fourfold degenerate sites: These are the third positions in codons that encode one of the two amino acids that are represented by four synonymous codons==.
For example, the amino acids alanine and proline are each encoded by four synonymous codons, so the third position in these codons is a fourfold degenerate site. - ==Twofold degenerate sites: These are the third positions in codons that encode one of the two amino acids that are represented by two synonymous codons==.
For example, the amino acids leucine, arginine, and serine each have two synonymous codons, so the third position in these codons is a twofold degenerate site. - ==Non-degenerate sites: These are the first and second positions in codons, which encode a specific amino acid without degeneracy==.
- ==Fourfold degenerate sites: These are the third positions in codons that encode one of the two amino acids that are represented by four synonymous codons==.
- Within a codon, there are three nucleotide positions.
- How Natural Selection creates Constraints, and on which particular Proteins?
- ==Natural selection can create functional constraints on proteins by favoring specific amino acid sequences that confer a selective advantage for a particular biological function==.
For example, in enzymes, certain amino acids may be crucial for catalytic activity, and substitutions that affect these residues could impair enzyme function.
Similarly, in proteins involved in protein-protein interactions, substitutions that disrupt key residues involved in the interaction could reduce protein function.
As a result, natural selection can act to preserve specific amino acid sequences in these functional regions, leading to the conservation of specific sites and the creation of functional constraints. - These constraints can be particularly strong in highly conserved proteins that perform critical functions across a wide range of organisms.
==For example, proteins involved in DNA replication or transcription are often highly conserved==, with substitutions at critical residues having severe consequences for protein function.
Conversely, proteins with less critical functions may be subject to weaker constraints, allowing for greater variability in sequence.
- ==Natural selection can create functional constraints on proteins by favoring specific amino acid sequences that confer a selective advantage for a particular biological function==.
—————————————————————
IMPORTANTE
IMPORTANTE Synonymus Substitutions: Nei casi in cui un cambiamento nucelotidico non varia la sequenza amminoacidica tradotta.
IMPORTANTE Non-Synonymus Substitutions: Un cambiamento nucelotidico che varierà la sequenza amminoacidica tradotta.
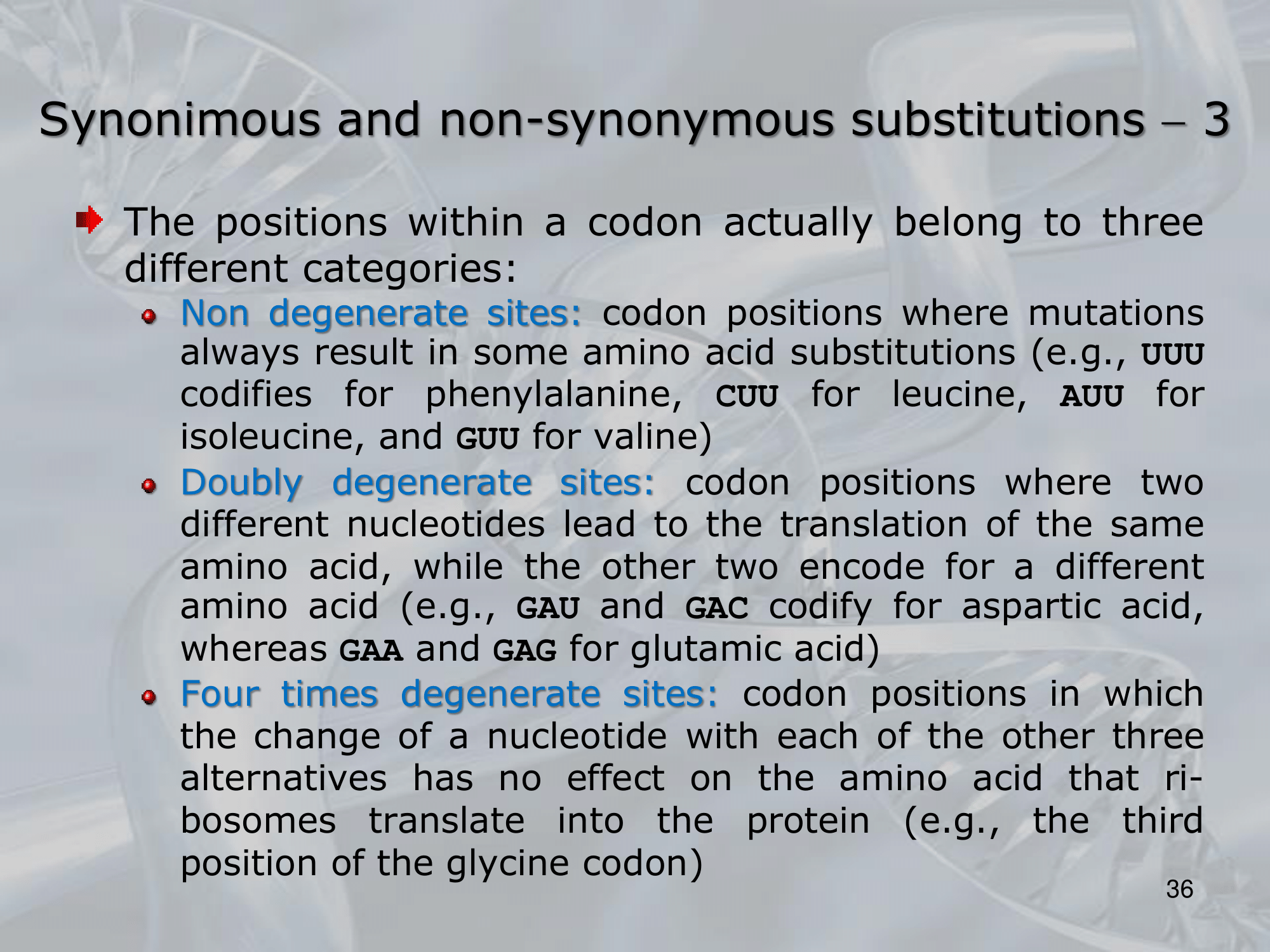
IMPORTANTE In un singolo amminoacido le sue basi possono essere divise in tre gruppi:
- Non Degenerate Sites: Posizione in cui qualsiasi cambiamente cambia l’amminoacido tradotto (~ex.: UUU ⇒ fenalanina, CUU ⇒ leucina, AUU ⇒ isoleucina, GUU ⇒ valina)
- Doubly Degenerate Sites: Una posizione in cui due nucleotidi codificano in uno stesso amminoacido, e altri due in un altro (~ex.: GAU e GAC codificano in acido aspartico, mentre GAG e GAA codificano in acido glutamminico)
- Four Times Degenerate Sites: Qualsiasi cambiamenti in questo sito risulta nello stesso amminoacido (~ex.: GGG, GGA, GGU, GGC, codificheranno tutti in glicina)
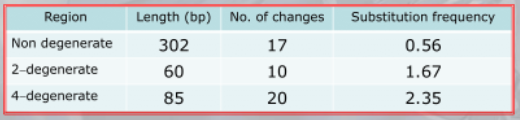
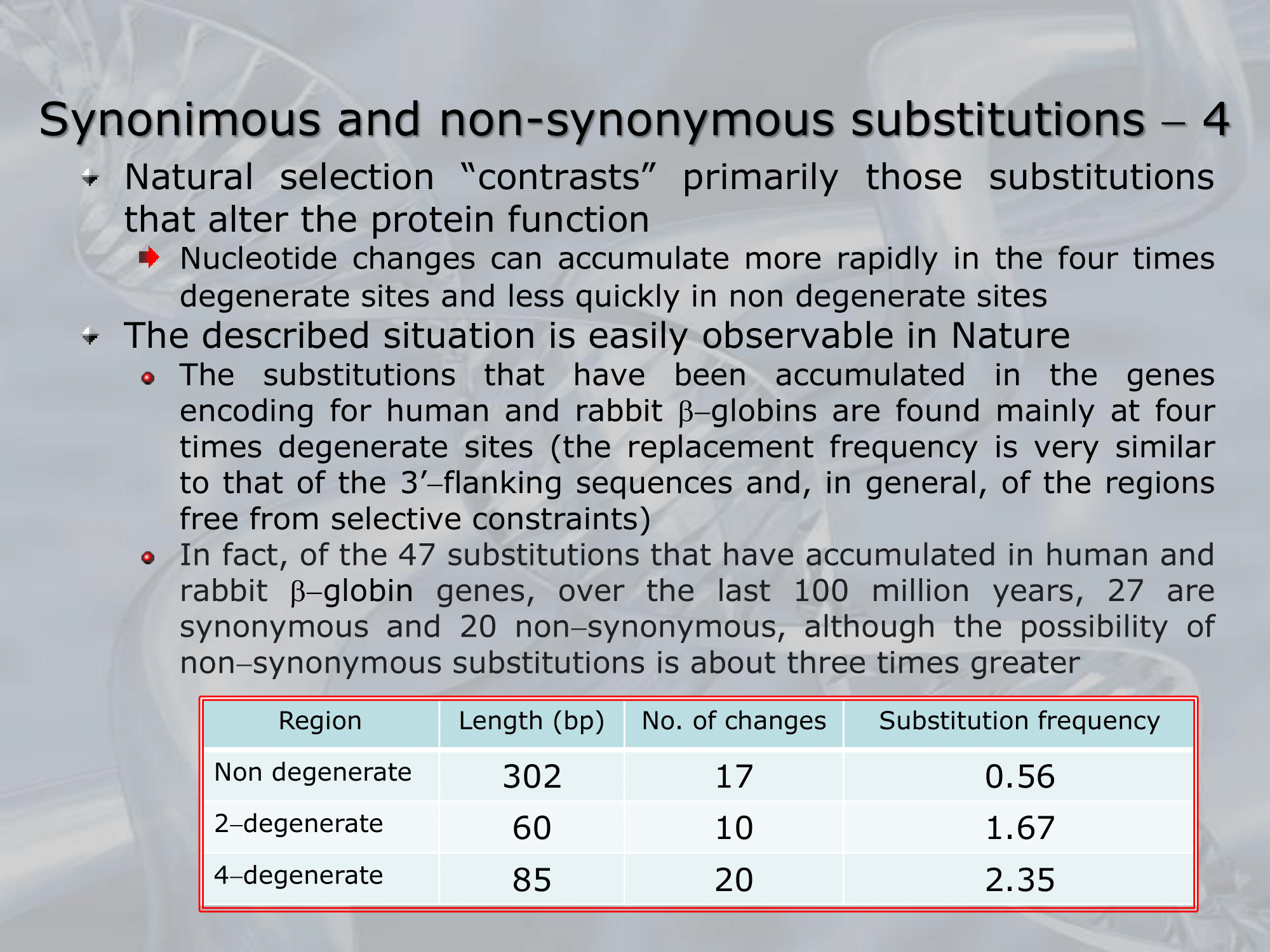
IMPORTANTE La selezione naturale tende a conservare le proteine che funzionano, infatti la frequenza di sostituzione che possiamo notare nella -globina nell’uomo è diversa per i tipi di siti:
—————————————————————
Slides with Notes




IMPORTANTE Synonymus Substitutions: Nei casi in cui un cambiamento nucelotidico non varia la sequenza amminoacidica tradotta.
IMPORTANTE Non-Synonymus Substitutions: Un cambiamento nucelotidico che varierà la sequenza amminoacidica tradotta.
IMPORTANTE In un singolo amminoacidi le sue basi possono essere divise in tre gruppi:
- Non Degenerate Sites: Posizione in cui qualsiasi cambiamente cambia l’amminoacido tradotto (~ex.: UUU ⇒ fenalanina, CUU ⇒ leucina, AUU ⇒ isoleucina, GUU ⇒ valina)
- Doubly Degenerate Sites: Una posizione in cui due nucleotidi codificano in uno stesso amminoacido, e altri due in un altro (~ex.: GAU e GAC codificano in acido aspartico, mentre GAG e GAA codificano in acido glutamminico)
- Four Times Degenerate Sites: Qualsiasi cambiamenti in questo sito risulta nello stesso amminoacido (~ex.: GGG, GGA, GGU, GGC, codificheranno tutti in glicina)
IMPORTANTE La selezione naturale tende a conservare le proteine che funzionano, infatti la frequenza di sostituzione che possiamo notare nella -globina nell’uomo è diversa per i tipi di siti: