Questions
- What are Introns and Exons?
- ==Introns and exons are two types of nucleotide sequences that are found in eukaryotic genes==.
They are part of the gene structure, and together they make up the protein-coding regions of a gene.- Exons are the parts of a gene that code for protein sequences. These regions are transcribed into RNA and then translated into amino acid sequences to form proteins.
Exons are usually found in multiple segments, with non-coding introns in between. The exact number and length of exons can vary between different genes and organisms. - Introns, on the other hand, are non-coding regions of DNA that are transcribed into RNA but are removed during the RNA processing step called splicing.
Introns are usually located between exons and do not code for protein sequences.
Introns can be quite long and complex, and their sequences can vary widely between different genes and organisms.
- Exons are the parts of a gene that code for protein sequences. These regions are transcribed into RNA and then translated into amino acid sequences to form proteins.
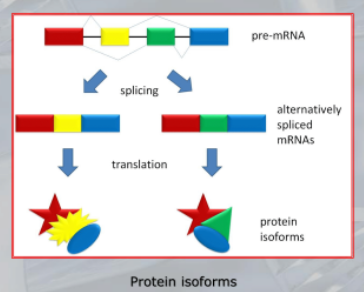
- The process of splicing removes the introns from the pre-mRNA transcript and joins together the exons to form the mature mRNA that is translated into protein.
This process is essential for generating diverse protein isoforms from a single gene and plays a critical role in regulating gene expression. - The presence of introns and exons in eukaryotic genes is a fundamental feature of the gene structure and contributes to the complexity and diversity of eukaryotic organisms.
The precise organization and arrangement of exons and introns can vary widely between genes and organisms and can have important implications for gene regulation, disease pathology, and evolutionary biology.
- ==Introns and exons are two types of nucleotide sequences that are found in eukaryotic genes==.
—————————————————————
IMPORTANTE
IMPORTANTE Differences in Translation of Eukaryotic and Prokaryotic In Prokaryotes the genes are transcribed in mRNA. In Eukaryotes the genes are transcribed in hnRNA (heterogeneus RNA) and then are alternativily spliced* into mRNA.
IMPORTANTE Alternative Splicing A process unique to eukaryotes. After the transcription process, RNA polymerase creates the hnRNA, that is then spliced by the splicesome (an ensamble of proteins devoted to splicing) to create the mRNA that will then be translated. “Splicing” is just the removal of introns (non-coding regions of eukaryotic genes), and the agglomeration of exons (coding regions of eukaryotic genes). The alternative splicing is the process in which different spliceosome components and regulatory factors can influence which exons are included or excluded in the final mRNA transcript, leading to a variety of possible splice variants.
NOTE: The poly A tail is a mean for the ribosomes to stop translation
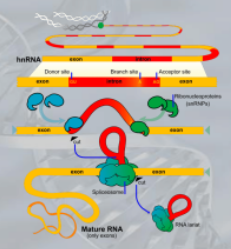
IMPORTANTE What are Introns? Many Eukaryotes genes contain sequences called introns that interrupt the coding regions. An Intron is a non-coding sequence of nucelotides that will not be present in the mature RNA. Introns are discarded. ~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
IMPORTANTE What are Exons? If we consider an eukaryotic gene, there might be some introns mixed within it, these exons wil be discarded, and what rimains, the actual coding region are called exons. ==Exons are NOT discarded==. ~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
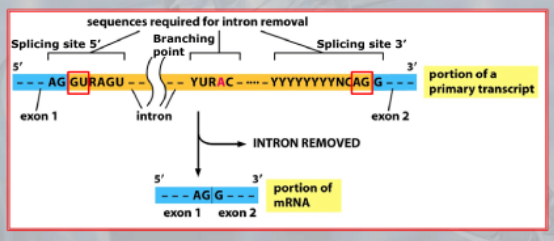
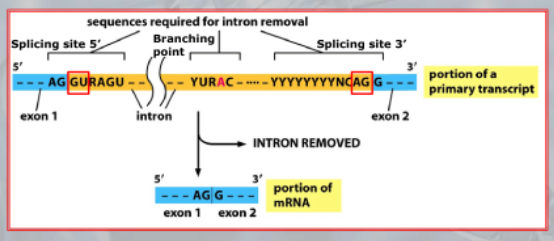
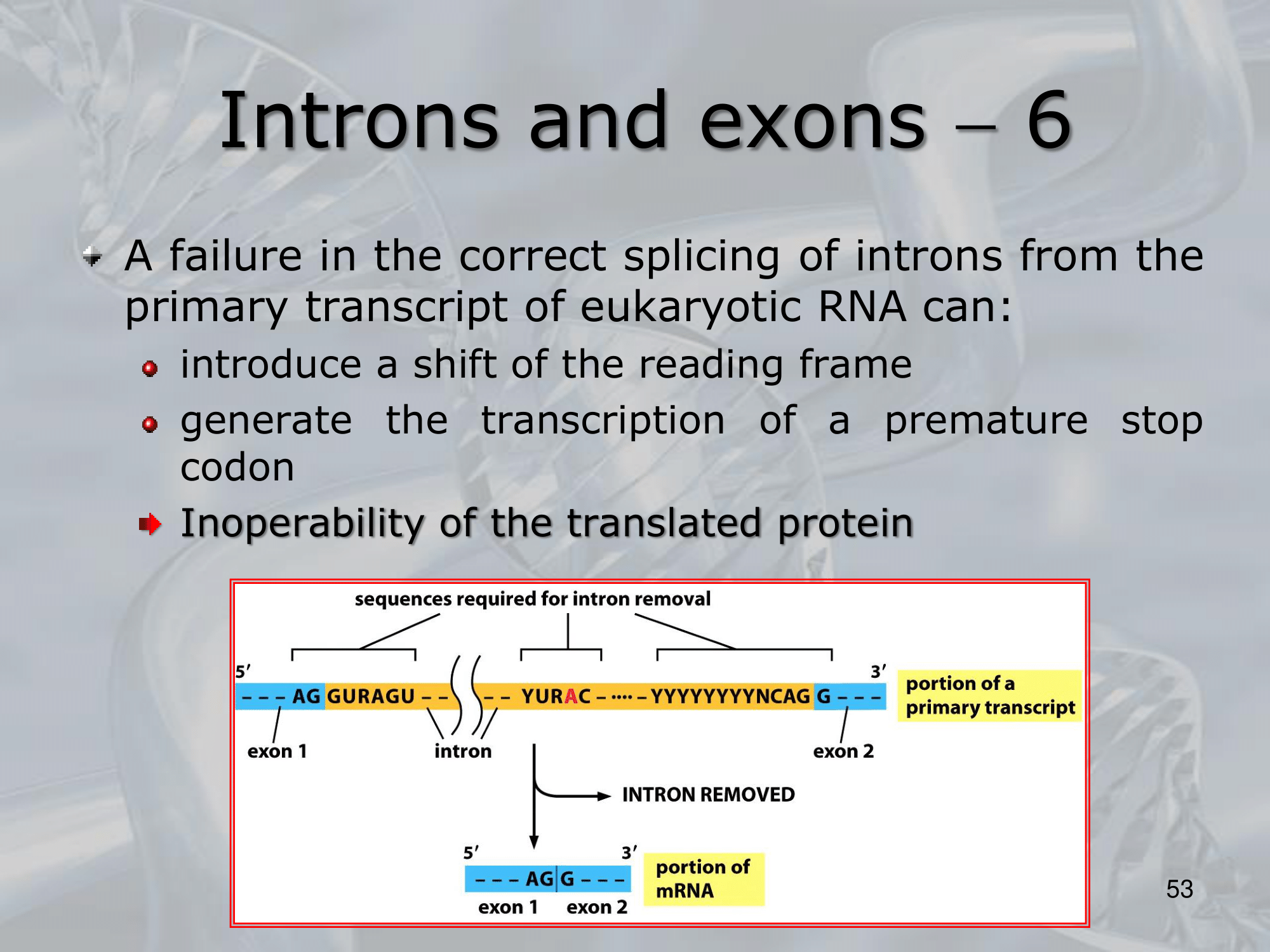
IMPORTANTE The GU-AG rule A rule used to recognize a specific type of intron in eukaryotic genes:
- The first pair sequence of nucleotides of this intron, is GU
- The last pair of nucleotides is AG.
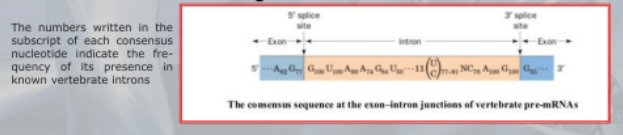
This AU-GU nucleotides (and others that appears in the middle of the introns sequence but less frequent than the AU-GU) are used as “markers” by the splicing apparatus when it will remove this intron. Most of the information on where the introns are found (to be deleted) are found in the introns itselves. Like a message saying “I’m an intron, delete me”
~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
IMPORTANTE Some statistics on Introns and Exons Introns have a minimum leght of 60 nucleotides, but no upper limit. Exons constitute about of our human genome, Introns about the . In of human genses there is at least one Intron.
IMPORTANTE Study on the position of Introns in genes: The length of introns and their nucleotide composition appear to be subject to weak selective constraint (it can change a lot during evolution, high mutation frequency) While the position of of the introns in a gene appears to be conserved from an evolutionary point of view. ==The introns often occupy the same position in homologous genes.==
Slides with Notes

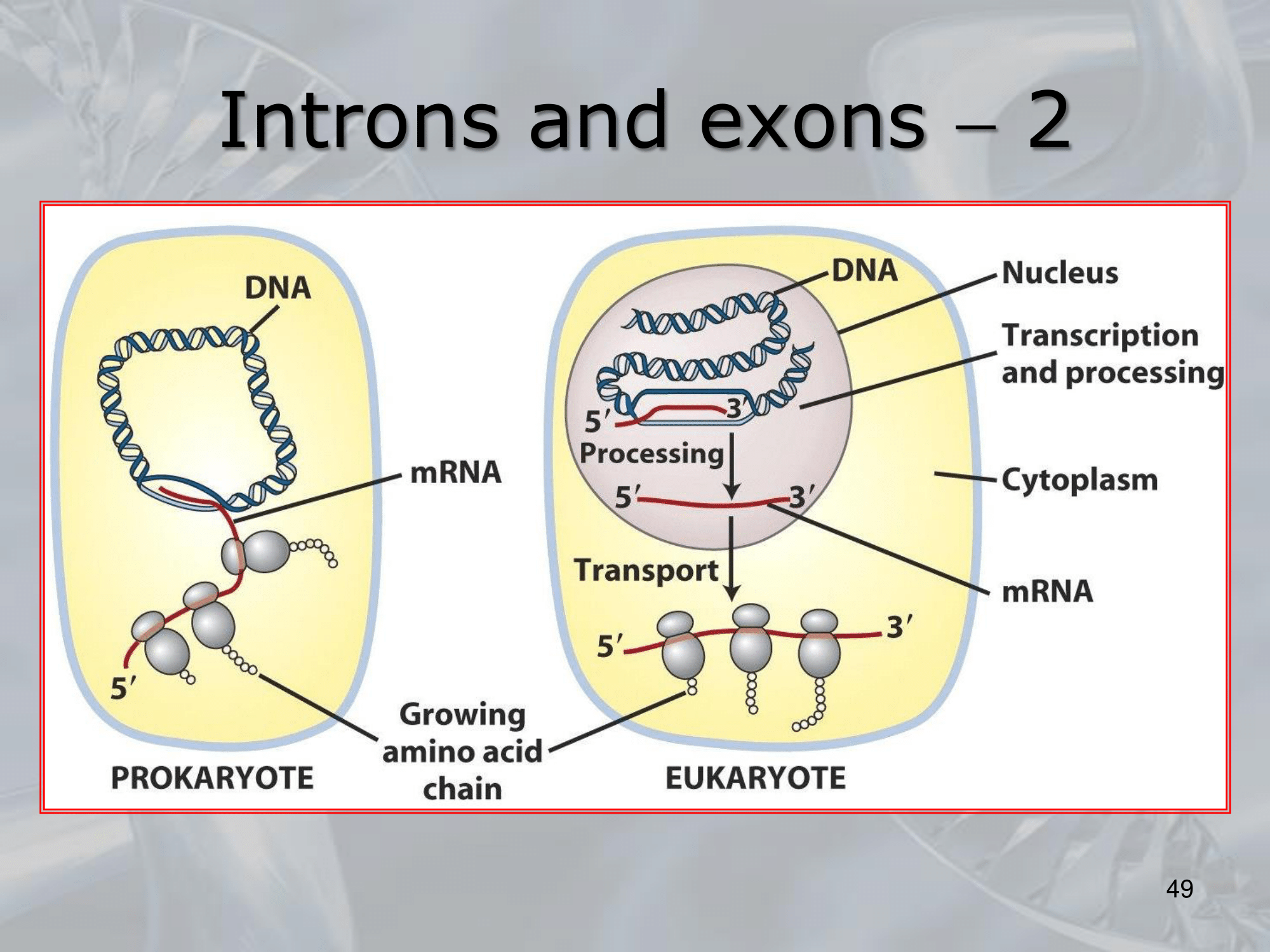
IMPORTANTE Differences in Translation of Eukaryotic and Prokaryotic In Prokaryotes the genes are transcribed in mRNA. In Eukaryotes the genes are transcribed in hnRNA (heterogeneus RNA) and then are alternativily spliced* into mRNA.
IMPORTANTE Alternative Splicing A process unique to eukaryotes. After the transcription process, RNA polymerase creates the hnRNA, that is then spliced by the splicesome (an ensamble of proteins devoted to splicing) to create the mRNA that will then be translated. “Splicing” is just the removal of introns (non-coding regions of eukaryotic genes), and the agglomeration of exons (coding regions of eukaryotic genes). The alternative splicing is the process in which different spliceosome components and regulatory factors can influence which exons are included or excluded in the final mRNA transcript, leading to a variety of possible splice variants.
NOTE: The poly A tail is a mean for the ribosomes to stop translation
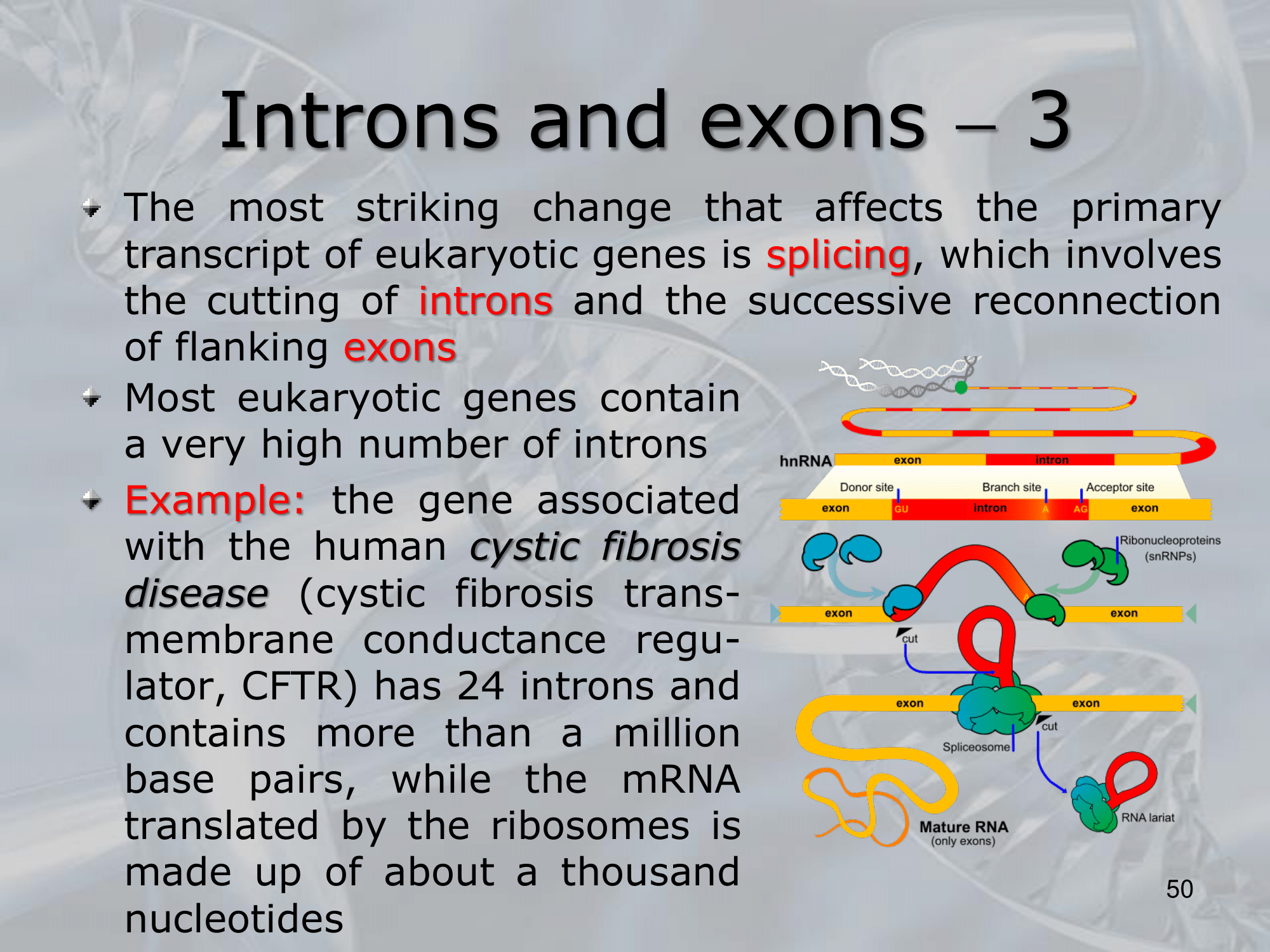
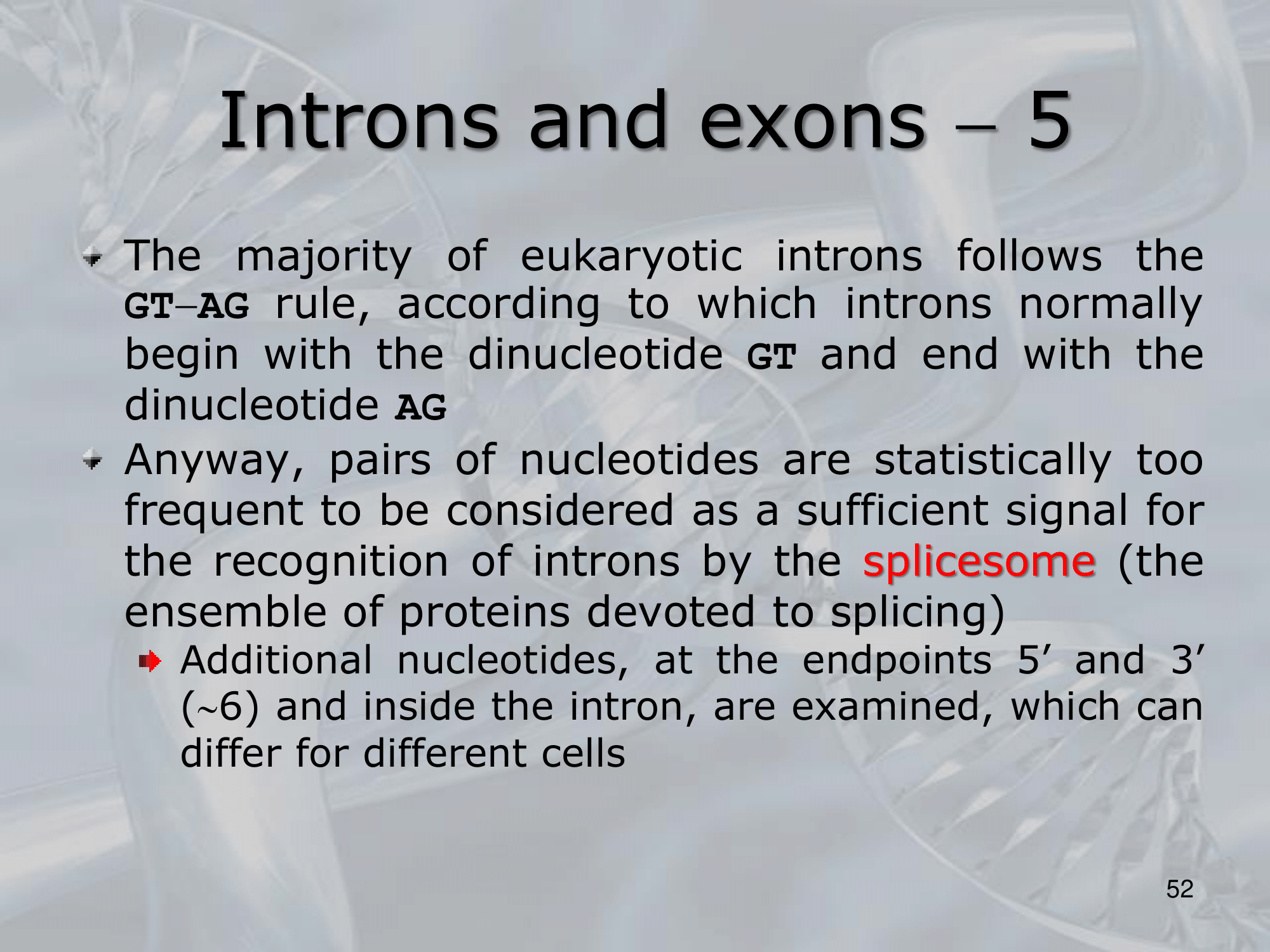
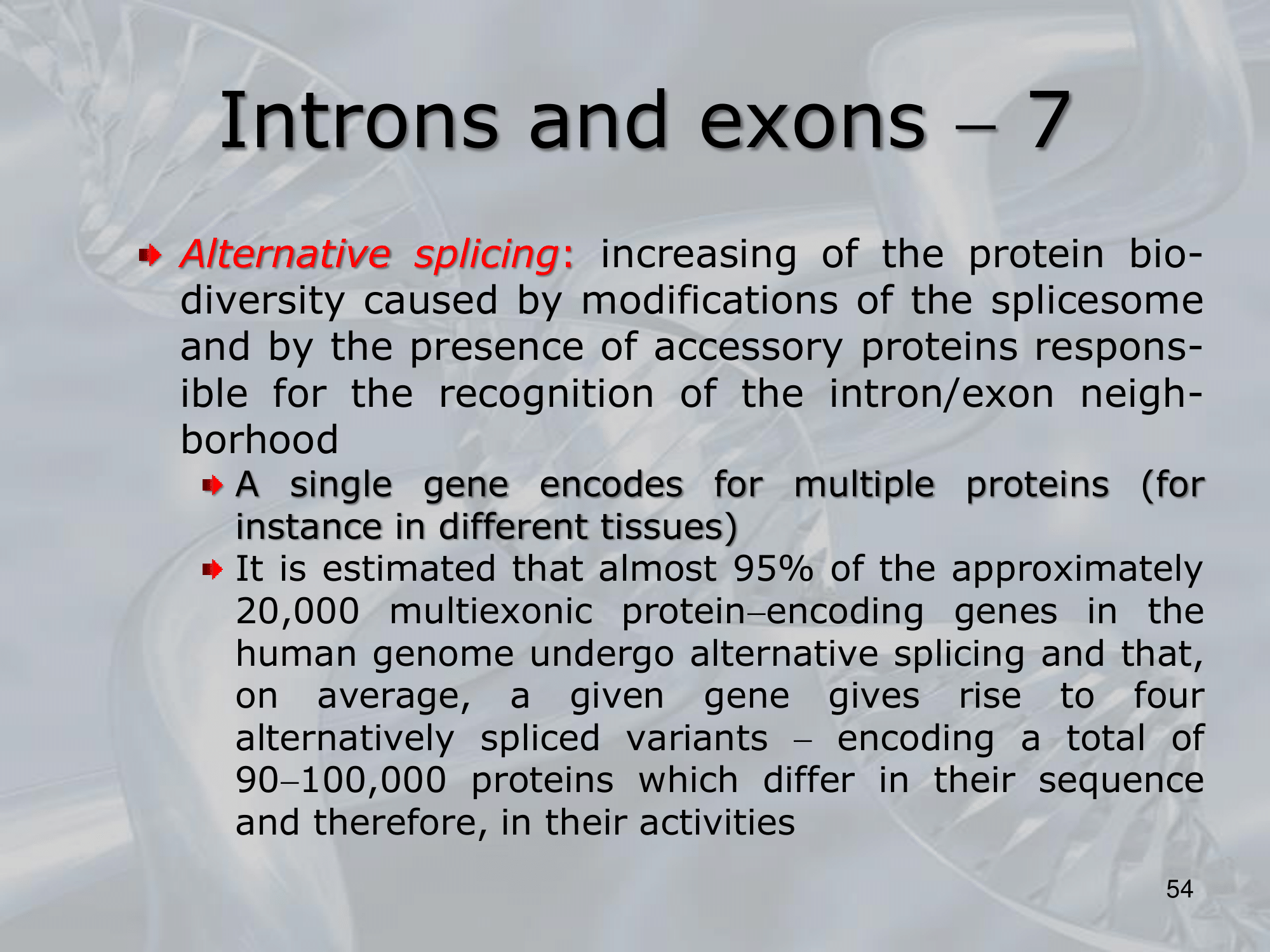
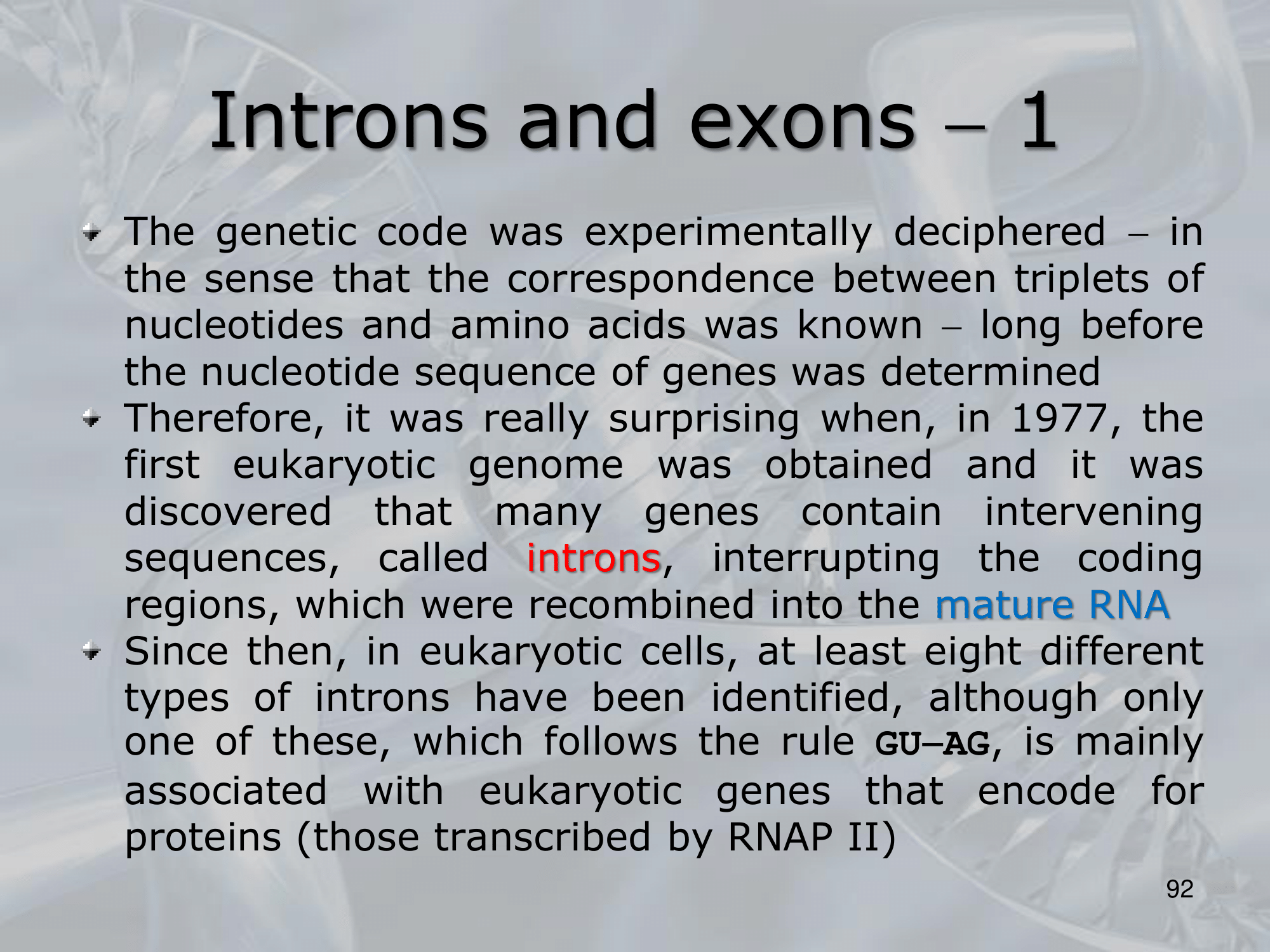


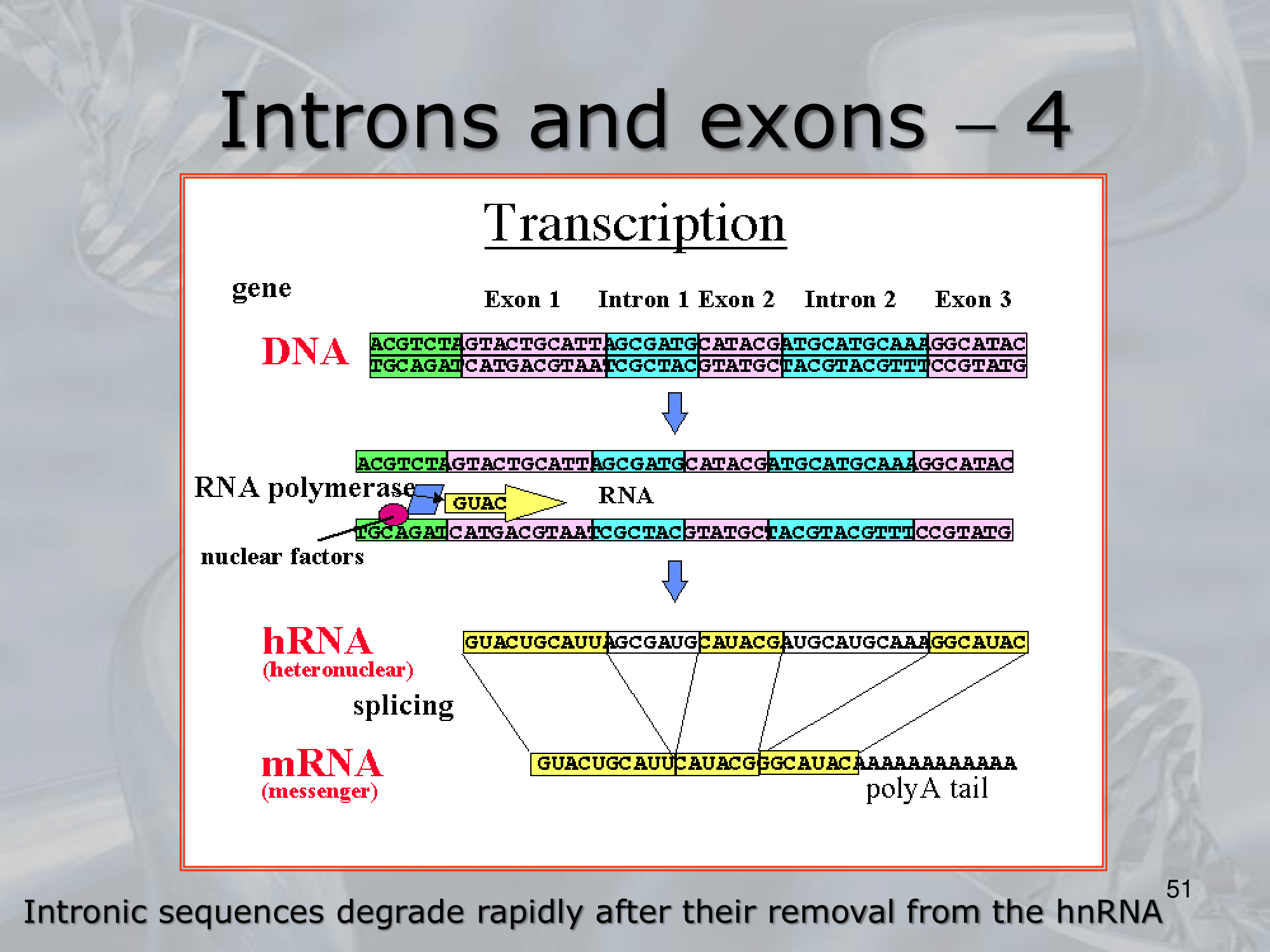


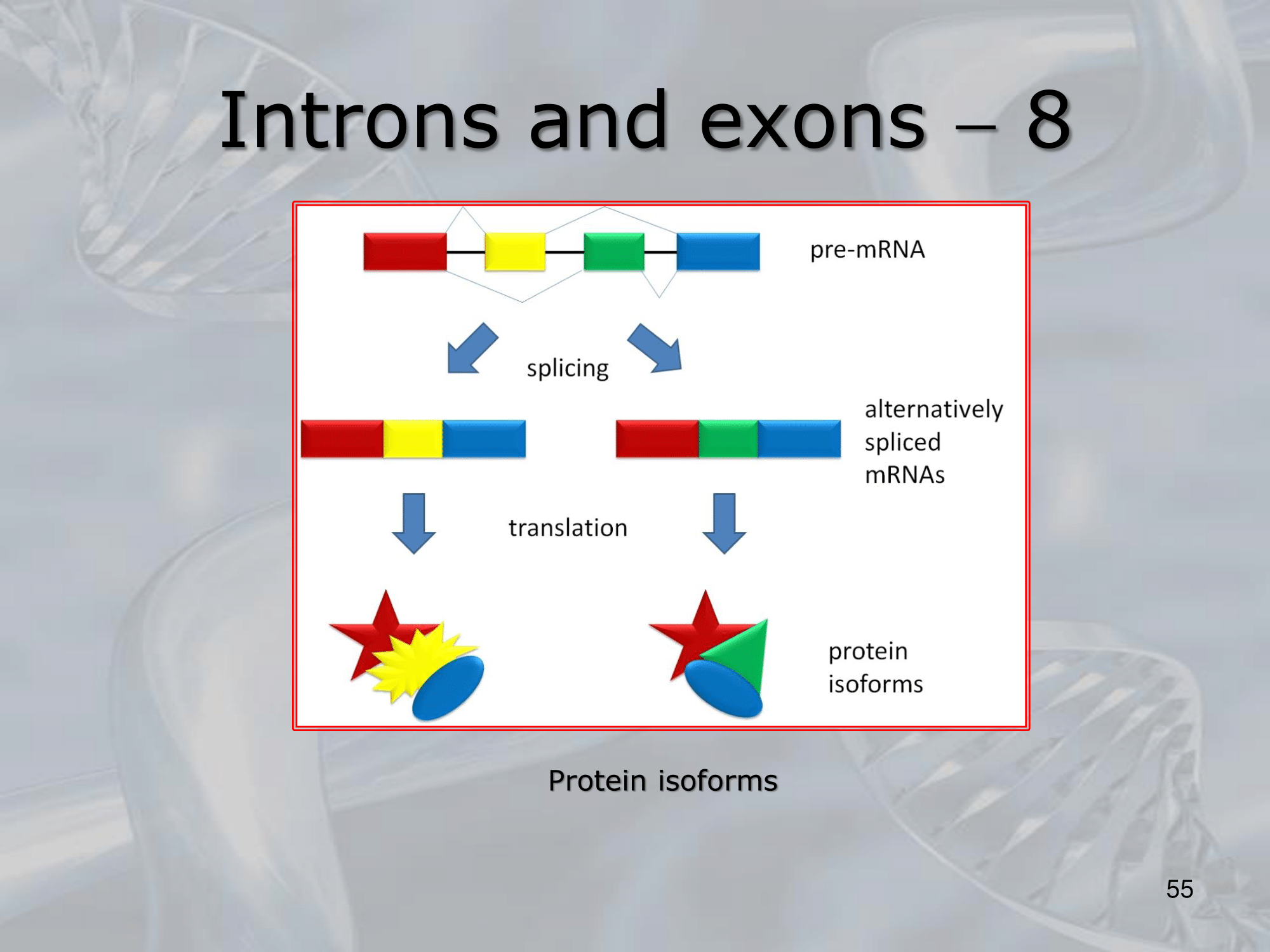

IMPORTANTE What are Introns? Many Eukaryotes genes contain sequences called introns that interrupt the coding regions. An Intron is a non-coding sequence of nucelotides that will not be present in the mature RNA. Introns are discarded. ~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
IMPORTANTE What are Exons? If we consider an eukaryotic gene, there might be some introns mixed within it, these exons wil be discarded, and what rimains, the actual coding region are called exons. Exons are not discarded. ~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
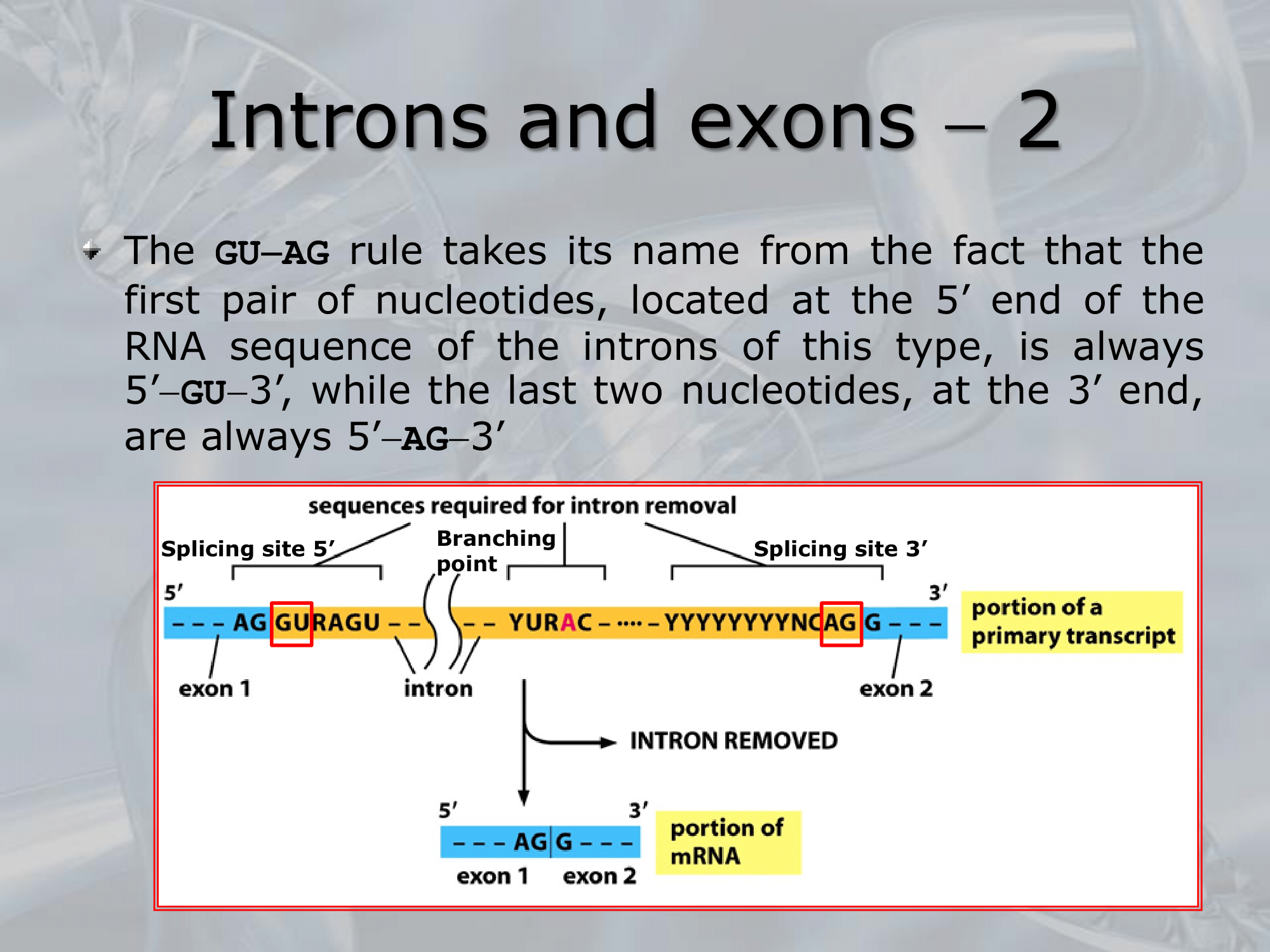
IMPORTANTE The GU-AG rule A rule used to recognize a specific type of intron in eukaryotic genes:
- The first pair sequence of nucleotides of this intron, is GU
- The last pair of nucleotides is AG.
This AU-GU nucleotides (and others that appears in the middle of the introns sequence but less frequent than the AU-GU) are used as “markers” by the splicing apparatus when it will remove this intron. Most of the information on where the introns are found (to be deleted) are found in the introns itselves. Like a message saying “I’m an intron, delete me”
~Ex.: of the GU-AG rule identifying an introns, and how the exons are then recombined together
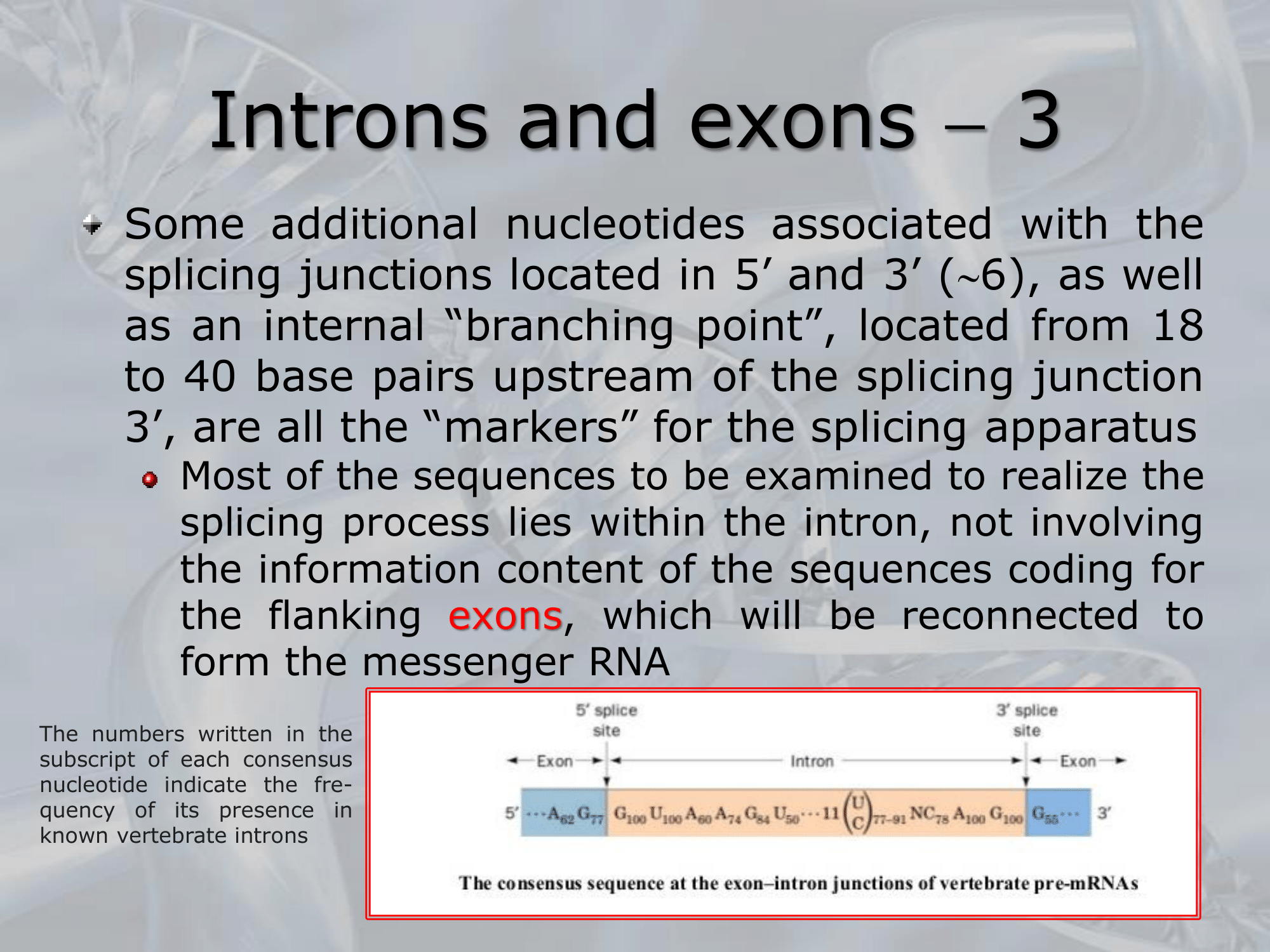


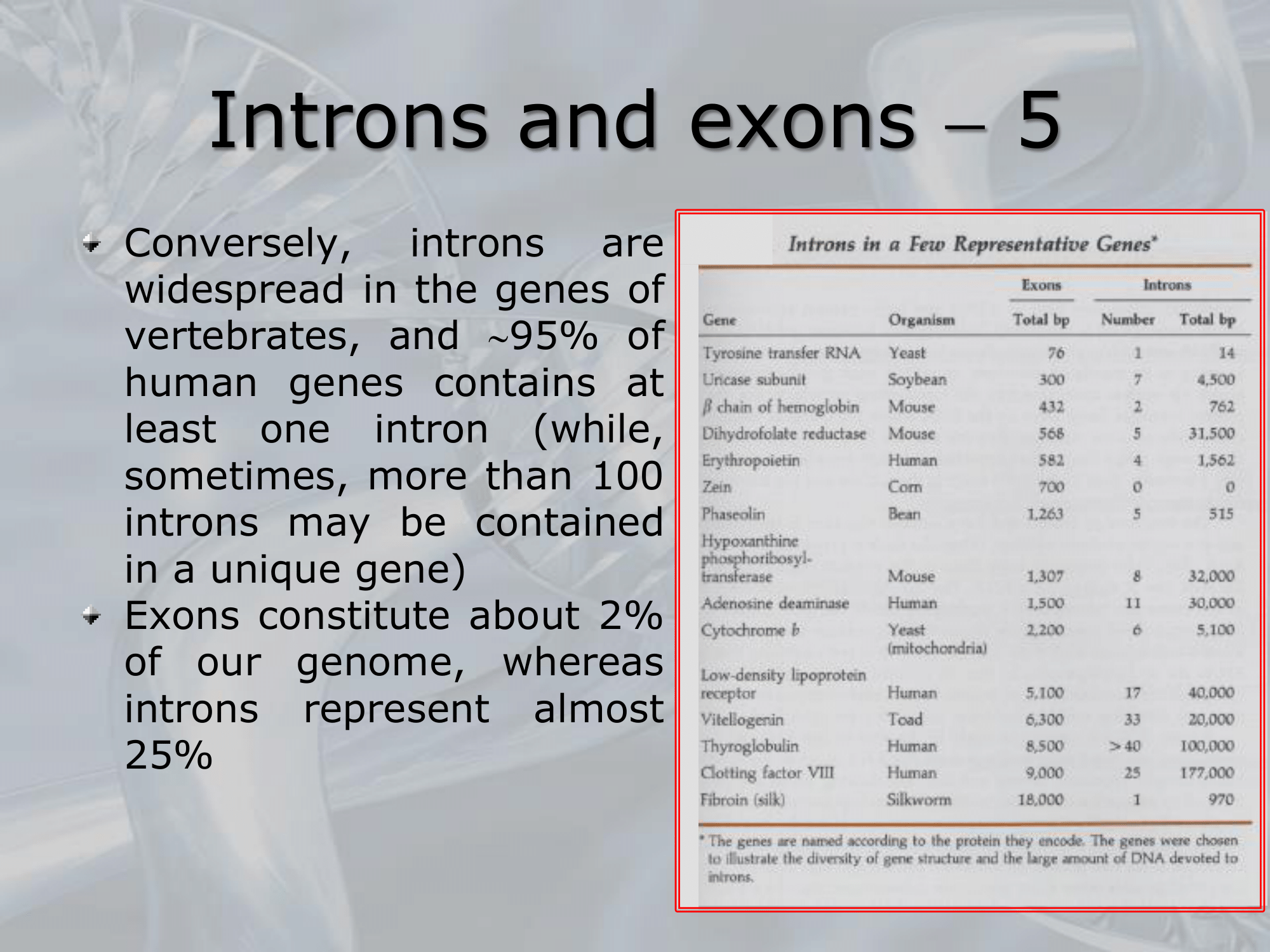
IMPORTANTE Some statistics on Introns and Exons Introns have a minimum leght of 60 nucleotides, but no upper limit. Exons constitute about of our human genome, Introns about the . In of human genses there is at least one Intron.

IMPORTANTE Study on the position of Introns in genes: The length of introns and their nucleotide composition appear to be subject to weak selective constraint (it can change a lot during evolution, high mutation frequency) While the position of of the introns in a gene appears to be conserved from an evolutionary point of view. ==The introns often occupy the same position in homologous genes.==