Questions
- What is an Open Reading Frame (ORF)?
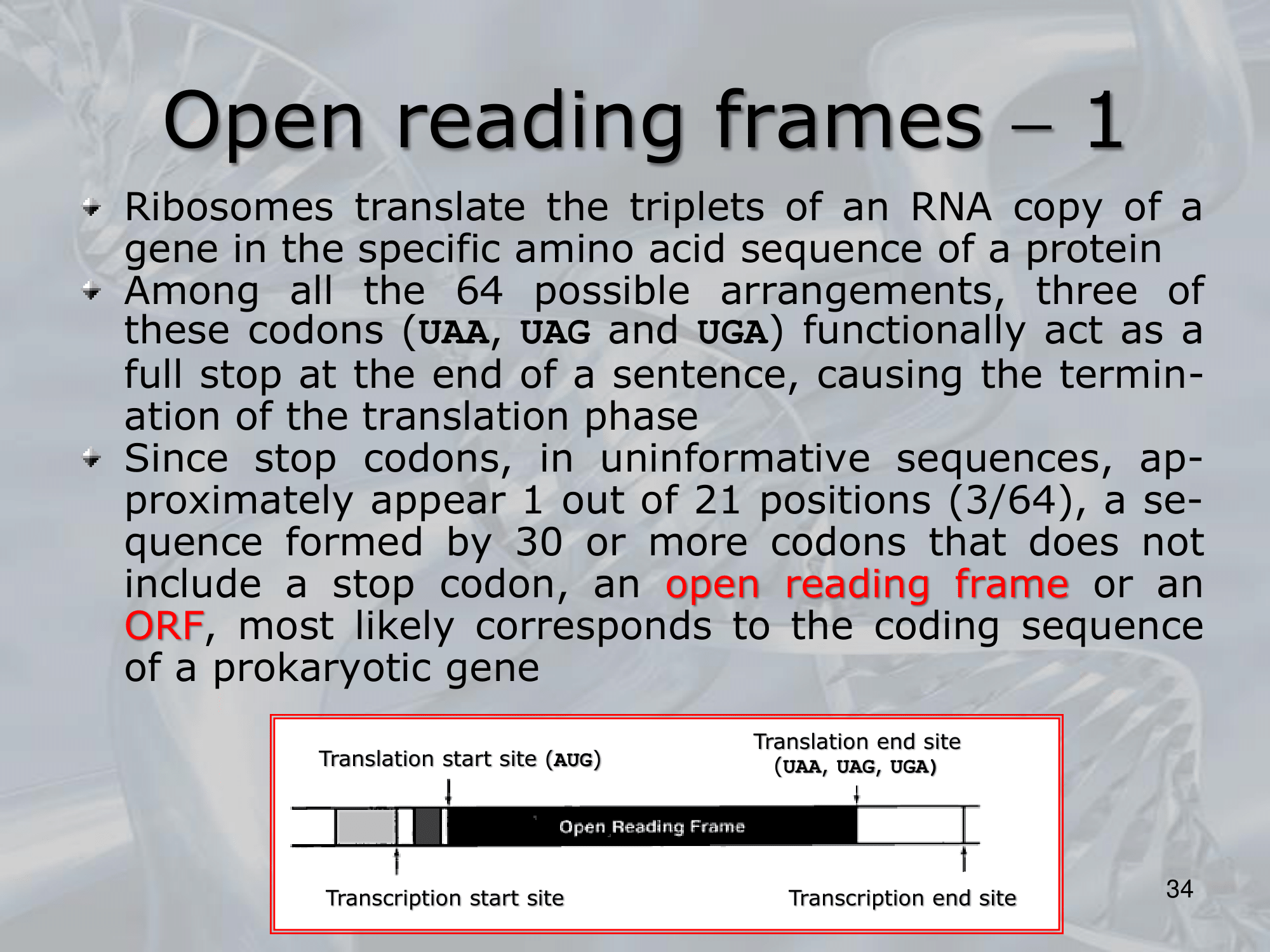
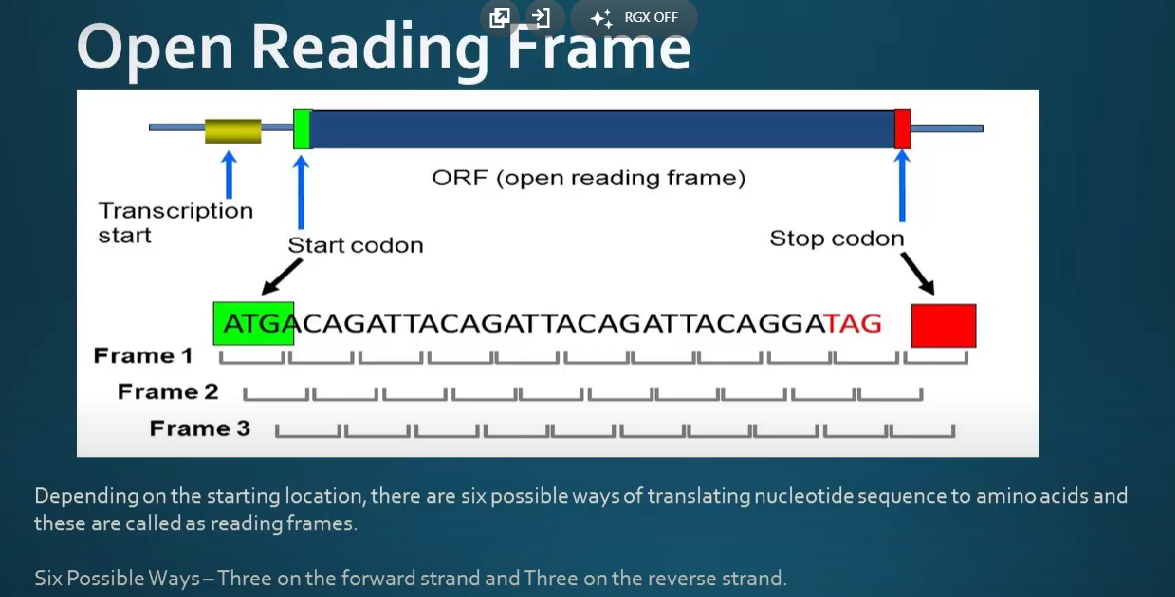
- An open reading frame (ORF) is a nucleotide sequence within a DNA or RNA molecule that has the potential to encode a protein.
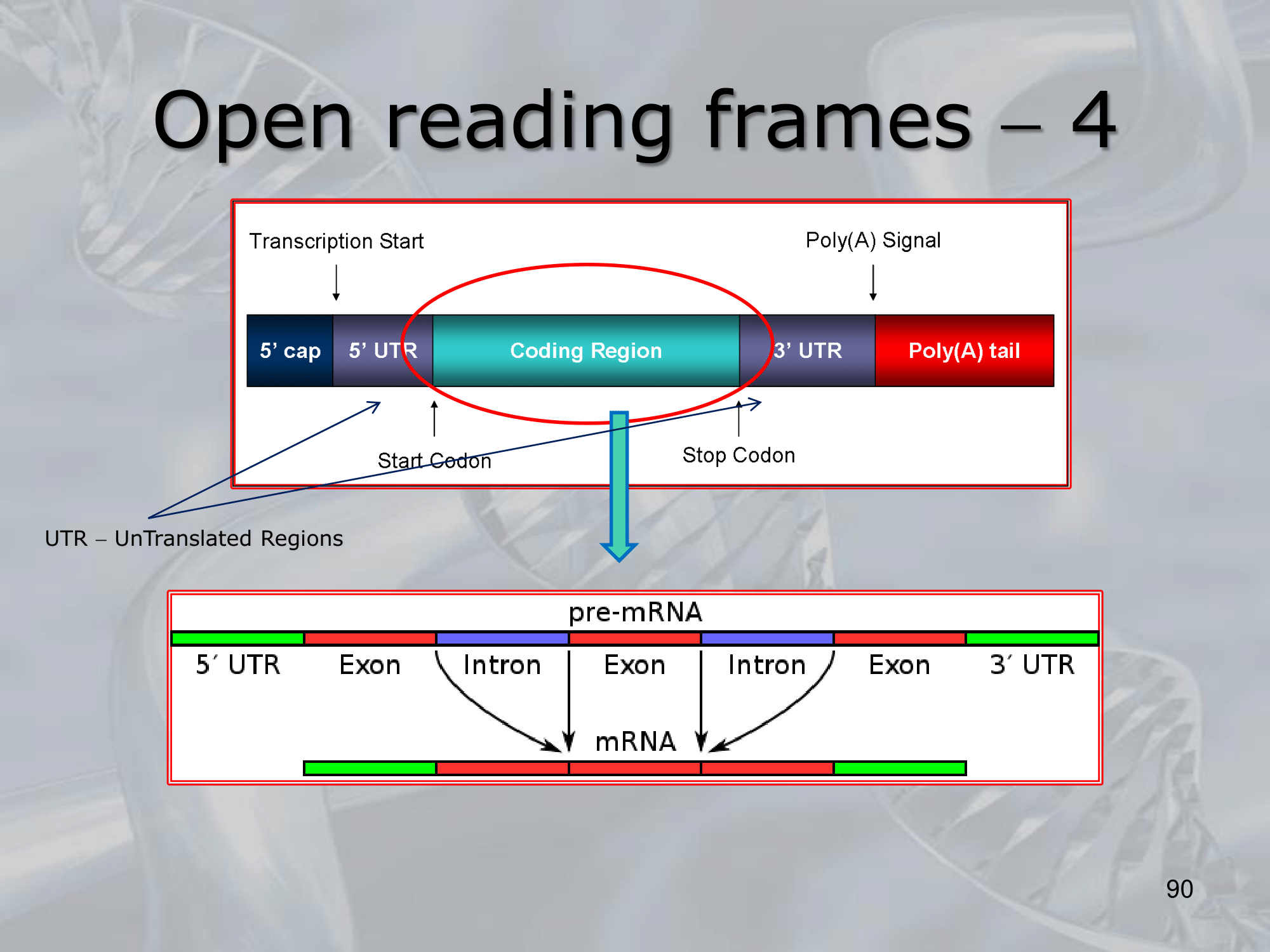
==An ORF begins with a start codon, which is typically AUG, and ends with a stop codon, which is typically UAA, UAG, or UGA==. - The term “open” refers to the fact that the sequence is not interrupted by any stop codons in the same reading frame, meaning that the sequence has the potential to be translated into a continuous polypeptide chain.
ORFs can be found in both coding and non-coding regions of DNA or RNA. - The identification and analysis of ORFs is an important step in genome annotation, and is often used to predict potential protein-coding genes within a genome sequence.
Bioinformatics tools can be used to scan DNA or RNA sequences for potential ORFs, and the resulting ORF predictions can be further analyzed to determine their function, expression, and potential role in cellular processes. - Overall, ORFs provide a useful framework for the analysis of potential protein-coding genes within a genome, and can be a valuable tool for understanding the genetic information encoded within a DNA or RNA sequence.
- An open reading frame (ORF) is a nucleotide sequence within a DNA or RNA molecule that has the potential to encode a protein.
—————————————————————
IMPORTANTE
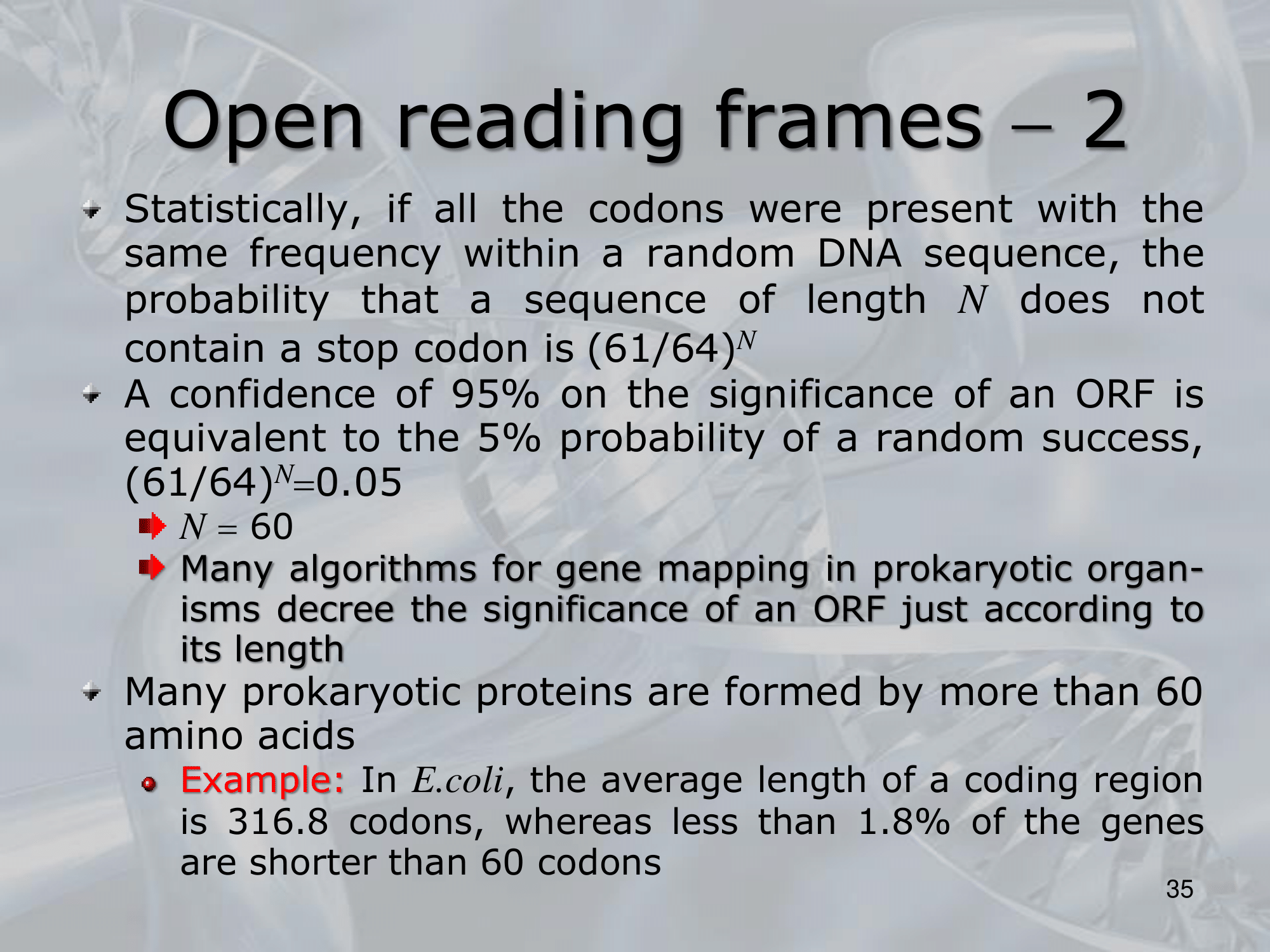
IMPORTANTE What is an ORF (Open Reading Frame)? An open reading frame, is a portion of a DNA sequence that does not include a stop codon (which functions as a stop signal). A long open reading frame is often part of a gene (that is, a sequence directly coding for a protein). Online Resource: National Human Genome Research Institute
IMPORTANTE What is a Codon? A codon is a DNA or RNA sequence of three nucleotides (a trinucleotide) that forms a unit of genomic information encoding a particular amino acid or signaling the termination of protein synthesis (stop codon). There are 64 different codons: 61 specify amino acids and 3 are used as stop codons. Online Resource: National Human Genome Research Institute
IMPORTANTE To form a single amino acid we need a triplet of an RNA copy of a gene
IMPORTANTE There exist 64 () possible arrangements to form each amino acid, but there exist only 20 different amino acids.
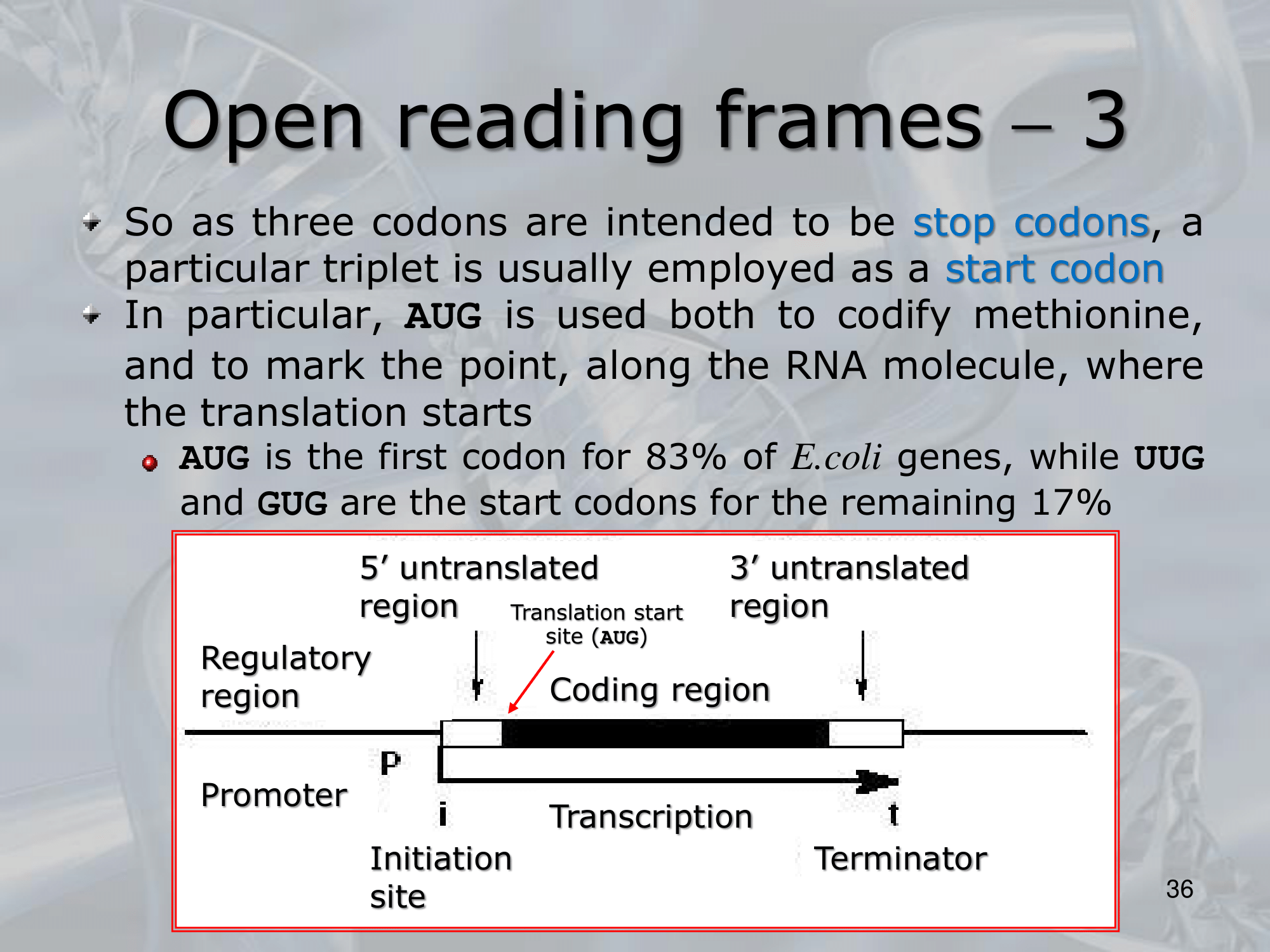
IMPORTANTE (In Prokaryotes) the start codon RNA code is AUG, UUG or GUG #IMPORTANTE (In Prokaryotes) the stop codon RNA code is UAA, UAG or UGA
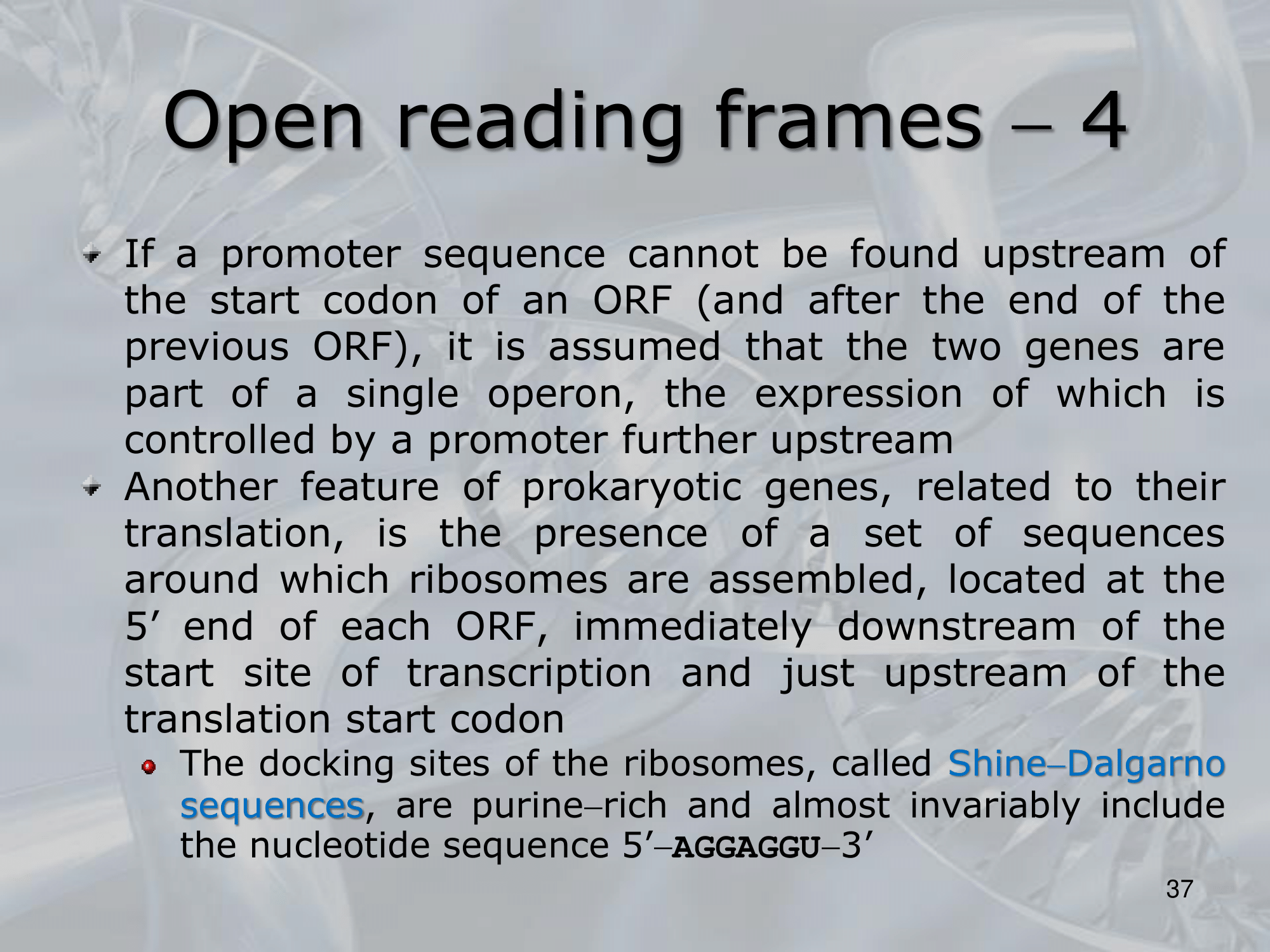
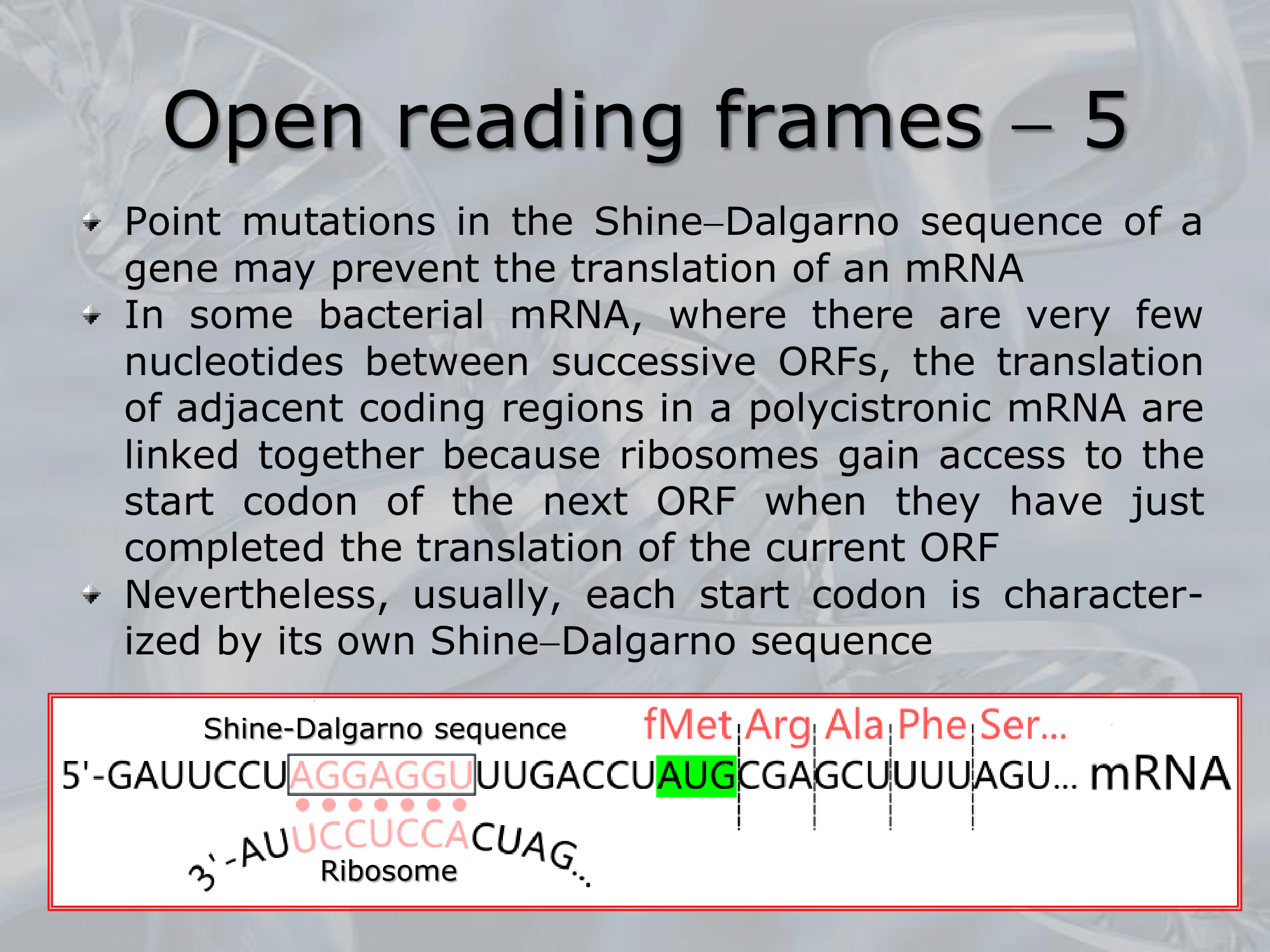
IMPORTANTE Shine-Dalgarno Sequence Most Prokaryotes genes have the so called Shine-Dalgarno sequence upstream the gene, it’s used by the ribosomes as a docking site.
IMPORTANTE In Prokaryotes if 2 genes are NOT separated by any promoter (neither one downstream the first, neither one upstream the second) we usually say that these two genes are part of the same operon
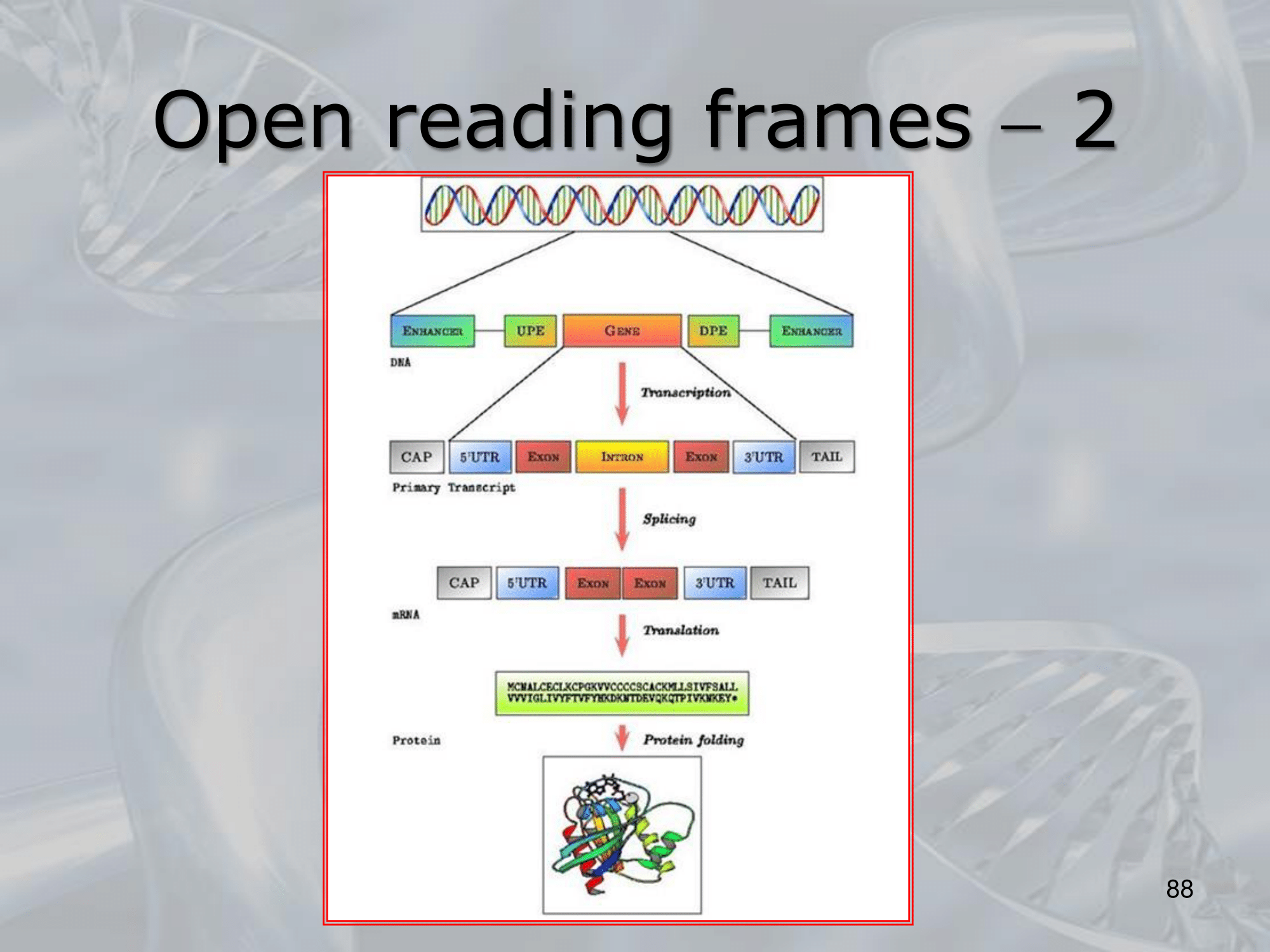
IMPORTANTE In Eukaryotes the DNA is first transcribed into hnRNA (heterogeneous nuclear RNA) which will undergo several changes, before being translated:
- Capping: “addition of a hood”, a set of chemical alteration always at the end of the 5’ end of the hnRNAs
- Splicing: “exon junction”, total and precise removal of sometimes very long segments inside the hnRNA
- Polyadenylation: “transforms hnRNA into mRNA”, replace the 3’ end of the hnRNA with a sequence of ~250 adenines (AA…AA)
Each of the three possible modification may occur differently types of cells
IMPORTANTE

Online Resource: Youtube
—————————————————————
Slides with Notes