Questions
- What is Comparative Modeling?
- ==Comparative modeling, also known as homology modeling, is a computational method for predicting the three-dimensional structure of a protein based on its amino acid sequence and the known structure of a homologous protein==.
==Homologous proteins are proteins that share significant sequence similarity with the protein of interest==, and their structures can be used as templates to predict the structure of the target protein. - Comparative modeling typically involves several steps.
- First, a suitable template protein with a known structure is identified based on sequence similarity.
- Next, the target protein sequence is aligned with the template protein sequence to identify corresponding positions of amino acids.
- Then, the alignment is used to generate a 3D model of the target protein structure based on the corresponding positions of amino acids in the template protein structure.
- The quality of the predicted model depends on the accuracy of the alignment between the target protein and the template protein and the quality of the template protein structure.
If the alignment is accurate and the template structure is high-quality, then the predicted model is likely to be accurate. However, if the alignment is inaccurate or the template structure is low-quality, then the predicted model may be less reliable. - Comparative modeling is a valuable tool for predicting the structure of proteins that are difficult to solve experimentally, such as membrane proteins or proteins that are too large or unstable to be crystallized.
It is also useful for predicting the structures of proteins in organisms where experimental methods for structure determination are not available.
- ==Comparative modeling, also known as homology modeling, is a computational method for predicting the three-dimensional structure of a protein based on its amino acid sequence and the known structure of a homologous protein==.
- What are exactly Protein Loops?
- ==Protein loops are regions in a protein structure that connect two secondary structural elements, (~ex. of secondary strucure elements: alpha helices or beta strands)==.
Loops are typically less structured than the secondary structural elements and can have variable lengths, conformations, and amino acid compositions. - Protein loops play an important role in protein function and stability, as they can contribute to protein-protein interactions, ligand binding, and enzymatic activity.
They can also be involved in protein folding and dynamics, as they are often more flexible than the surrounding secondary structural elements. - Due to their structural variability, protein loops can be difficult to predict accurately using computational methods.
However, understanding the properties of protein loops is important for understanding protein structure and function and for developing computational methods for predicting loop conformations.
- ==Protein loops are regions in a protein structure that connect two secondary structural elements, (~ex. of secondary strucure elements: alpha helices or beta strands)==.
—————————————————————
IMPORTANTE
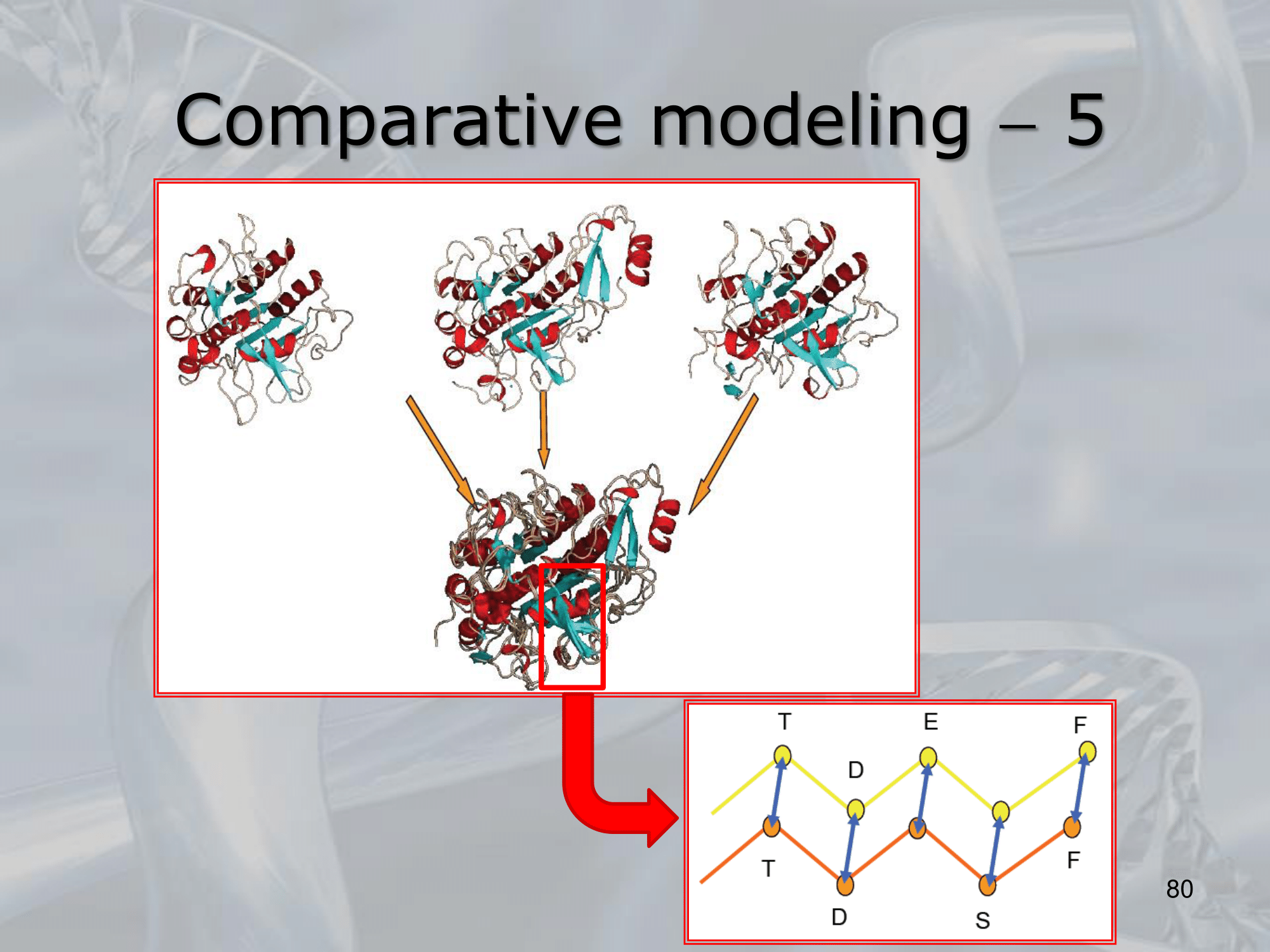
IMPORTANTE Comparative Modelling: Using similar already known proteins we create a not-yet-known 3D folding structure of a protein.
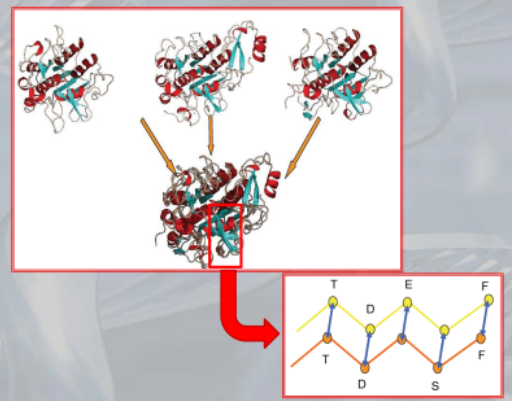
Algorithm:
- Identification of a set of protein structures realted to the target protein.
- Alignment of the target sequence with the template protein sequence
- Building the model: we overlap the loops (the backbone of the protein) and then using already known protein we build the loop and the sidechain
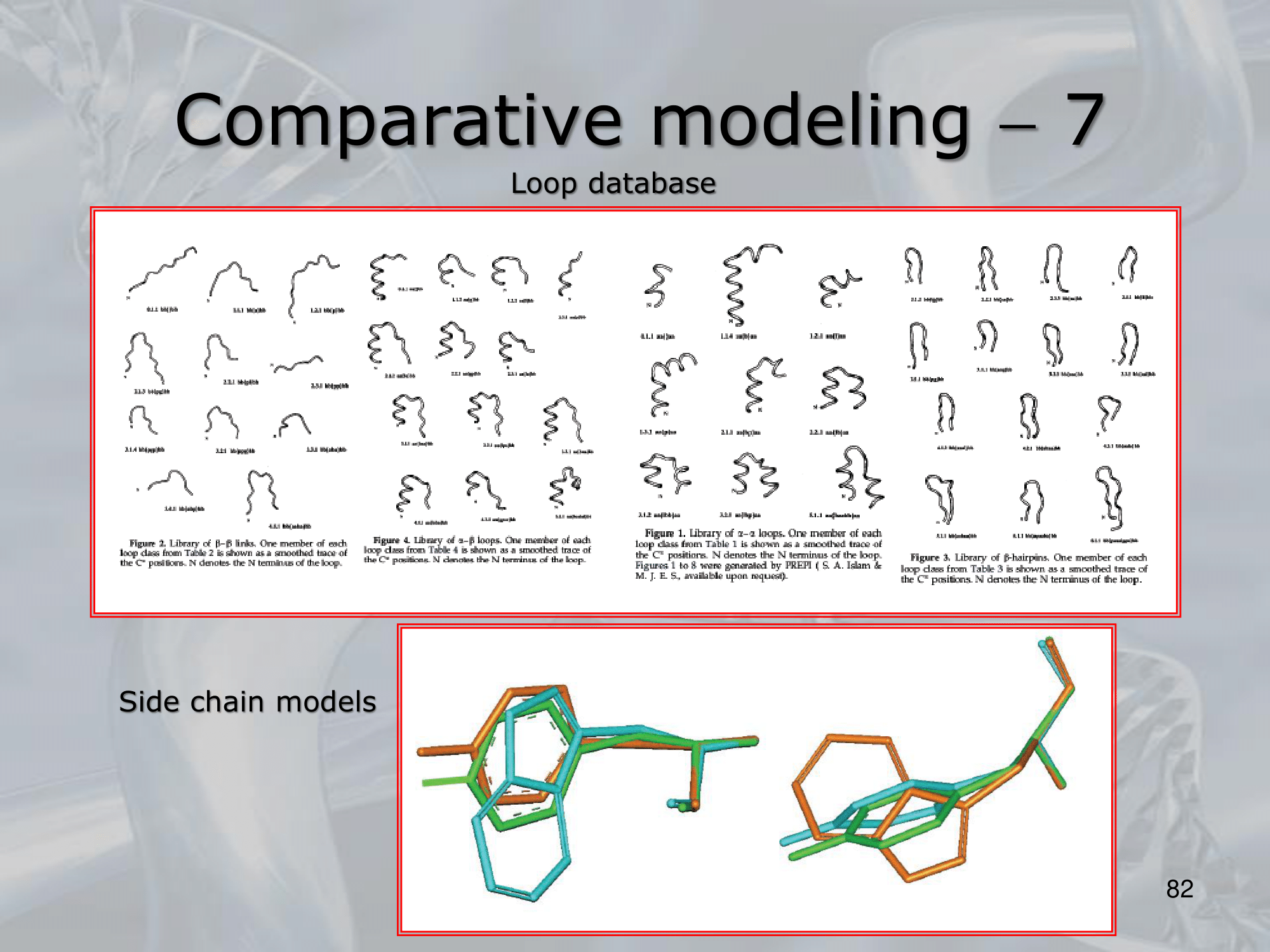
- Loop Modelling: using a database of already known loops.
- Side Chain Modelling: using rotaries libraries
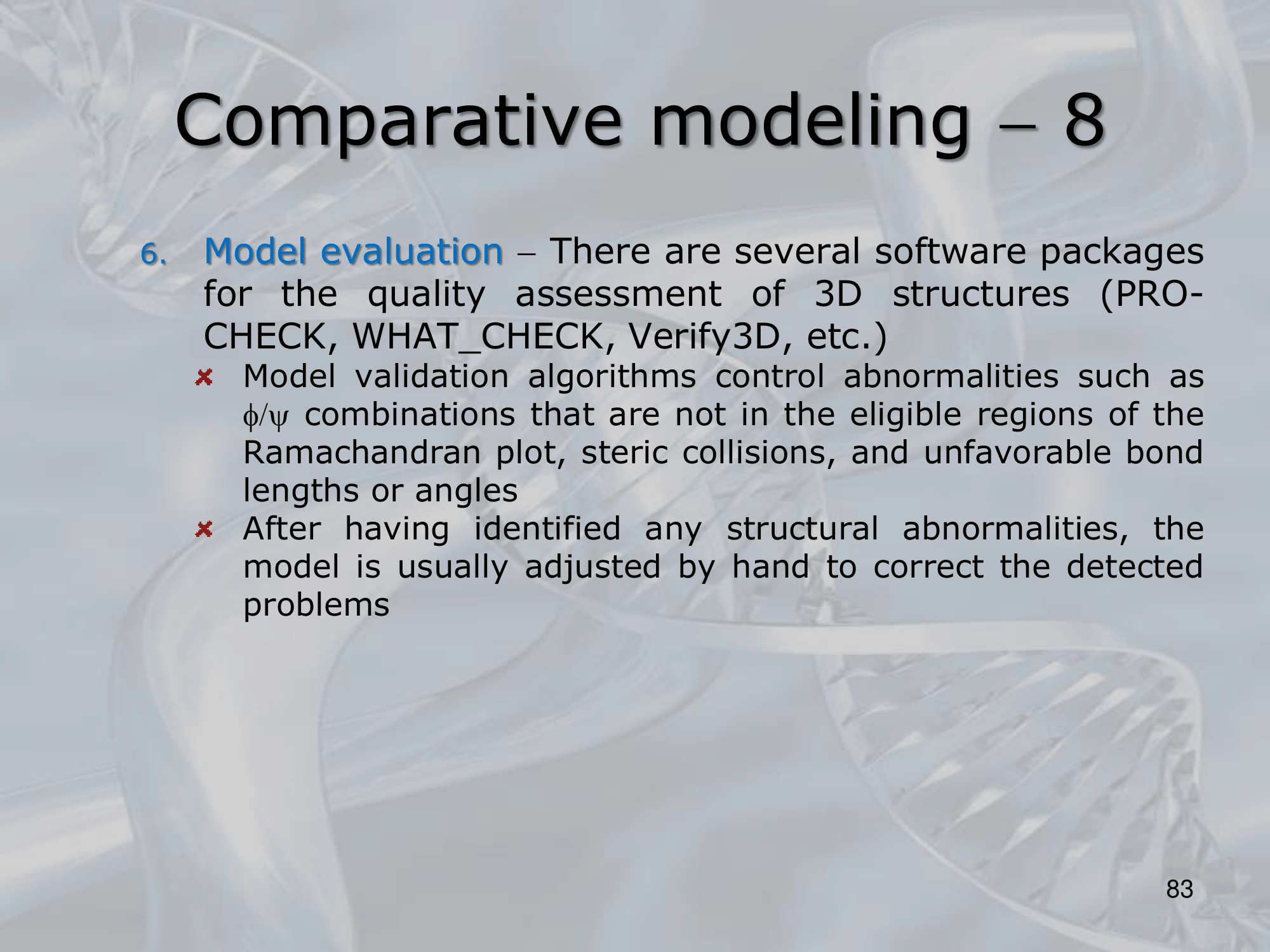
- Model Evaluation (using well known softwer packages: CHECK, WHAT_CHECK, Verify3D, )
—————————————————————
Slides with Notes

IMPORTANTE Comparative Modelling: Using similar already known proteins we create a not-yet-known 3D folding structure of a protein.

Algorithm:
- Identification of a set of protein structures realted to the target protein.
- Alignment of the target sequence with the template protein sequence
- Building the model: we overlap the loops (the backbone of the protein) and then using already known protein we build the loop and the sidechain
- Loop Modelling: using a database of already known loops.
- Side Chain Modelling: using rotaries libraries
- Model Evaluation (using well known softwer packages: CHECK, WHAT_CHECK, Verify3D, )
TODO What are exactly protein loops?